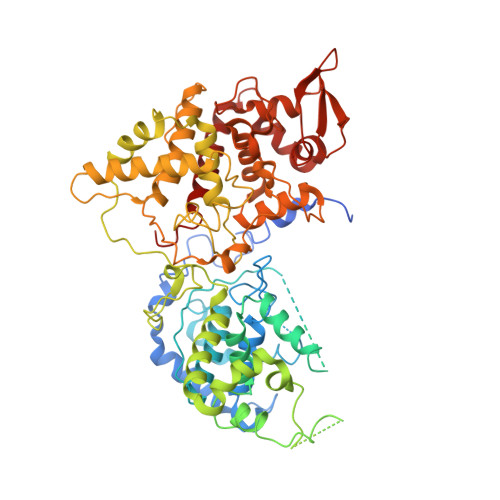
Using cryo-EM to understand antimycobacterial resistance in the catalase-peroxidase (KatG) from Mycobacterium tuberculosis.
Munir, A., Wilson, M.T., Hardwick, S.W., Chirgadze, D.Y., Worrall, J.A.R., Blundell, T.L., Chaplin, A.K.(2021) Structure 29: 899-912.e4
- PubMed: 33444527 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2020.12.008
- Primary Citation Related Structures:
7A2I, 7A7A, 7A7C, 7A8Z, 7AA3, 7AG8 - PubMed Abstract:
Resolution advances in cryoelectron microscopy (cryo-EM) now offer the possibility to visualize structural effects of naturally occurring resistance mutations in proteins and also of understanding the binding mechanisms of small drug molecules. In Mycobacterium tuberculosis the multifunctional heme enzyme KatG is indispensable for activation of isoniazid (INH), a first-line pro-drug for treatment of tuberculosis. We present a cryo-EM methodology for structural and functional characterization of KatG and INH resistance variants. The cryo-EM structure of the 161 kDa KatG dimer in the presence of INH is reported to 2.7 Å resolution allowing the observation of potential INH binding sites. In addition, cryo-EM structures of two INH resistance variants, identified from clinical isolates, W107R and T275P, are reported. In combination with electronic absorbance spectroscopy our cryo-EM approach reveals how these resistance variants cause disorder in the heme environment preventing heme uptake and retention, providing insight into INH resistance.
- Department of Biochemistry, University of Cambridge, Cambridge, CB2 1GA, UK.
Organizational Affiliation: