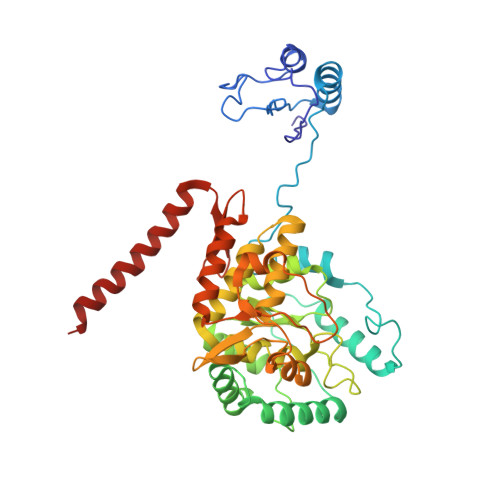
Structural mechanism for tyrosine hydroxylase inhibition by dopamine and reactivation by Ser40 phosphorylation.
Bueno-Carrasco, M.T., Cuellar, J., Flydal, M.I., Santiago, C., Krakenes, T.A., Kleppe, R., Lopez-Blanco, J.R., Marcilla, M., Teigen, K., Alvira, S., Chacon, P., Martinez, A., Valpuesta, J.M.(2022) Nat Commun 13: 74-74
- PubMed: 35013193 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-27657-y
- Primary Citation Related Structures:
6ZN2, 6ZVP, 6ZZU, 7A2G, 7PIM - PubMed Abstract:
Tyrosine hydroxylase (TH) catalyzes the rate-limiting step in the biosynthesis of dopamine (DA) and other catecholamines, and its dysfunction leads to DA deficiency and parkinsonisms. Inhibition by catecholamines and reactivation by S40 phosphorylation are key regulatory mechanisms of TH activity and conformational stability. We used Cryo-EM to determine the structures of full-length human TH without and with DA, and the structure of S40 phosphorylated TH, complemented with biophysical and biochemical characterizations and molecular dynamics simulations. TH presents a tetrameric structure with dimerized regulatory domains that are separated 15 Å from the catalytic domains. Upon DA binding, a 20-residue α-helix in the flexible N-terminal tail of the regulatory domain is fixed in the active site, blocking it, while S40-phosphorylation forces its egress. The structures reveal the molecular basis of the inhibitory and stabilizing effects of DA and its counteraction by S40-phosphorylation, key regulatory mechanisms for homeostasis of DA and TH.
- Centro Nacional de Biotecnología (CNB-CSIC), Madrid, Spain.
Organizational Affiliation: