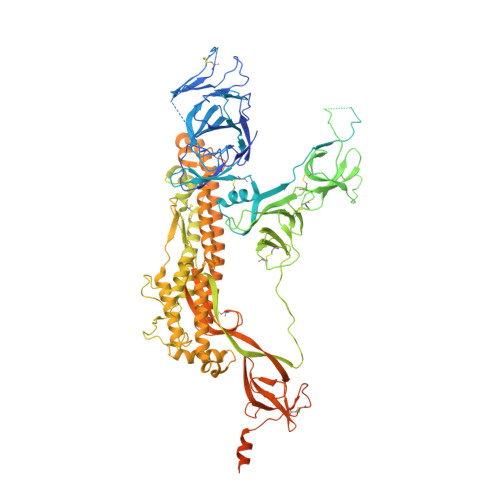
Continuous flexibility analysis of SARS-CoV-2 spike prefusion structures.
Melero, R., Sorzano, C.O.S., Foster, B., Vilas, J.L., Martinez, M., Marabini, R., Ramirez-Aportela, E., Sanchez-Garcia, R., Herreros, D., Del Cano, L., Losana, P., Fonseca-Reyna, Y.C., Conesa, P., Wrapp, D., Chacon, P., McLellan, J.S., Tagare, H.D., Carazo, J.M.(2020) IUCrJ 7: 1059-1069
- PubMed: 33063791 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2052252520012725
- Primary Citation Related Structures:
6ZOW, 6ZP5, 6ZP7 - PubMed Abstract:
Using a new consensus-based image-processing approach together with principal component analysis, the flexibility and conformational dynamics of the SARS-CoV-2 spike in the prefusion state have been analysed. These studies revealed concerted motions involving the receptor-binding domain (RBD), N-terminal domain, and subdomains 1 and 2 around the previously characterized 1-RBD-up state, which have been modeled as elastic deformations. It is shown that in this data set there are not well defined, stable spike conformations, but virtually a continuum of states. An ensemble map was obtained with minimum bias, from which the extremes of the change along the direction of maximal variance were modeled by flexible fitting. The results provide a warning of the potential image-processing classification instability of these complicated data sets, which has a direct impact on the interpretability of the results.
- Centro Nacional de Biotecnologia-CSIC, Calle Darwin 3, 28049 Cantoblanco, Madrid, Spain.
Organizational Affiliation: