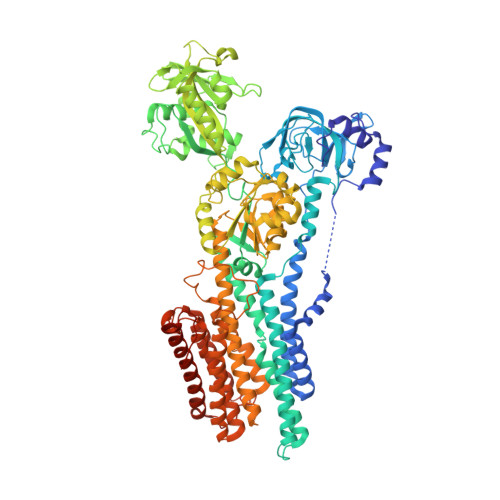
The Crystal Structure of the Ca 2+ -ATPase 1 from Listeria monocytogenes reveals a Pump Primed for Dephosphorylation.
Hansen, S.B., Dyla, M., Neumann, C., Quistgaard, E.M.H., Andersen, J.L., Kjaergaard, M., Nissen, P.(2021) J Mol Biology 433: 167015-167015
- PubMed: 33933469 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2021.167015
- Primary Citation Related Structures:
6ZHF, 6ZHG, 6ZHH - PubMed Abstract:
Many bacteria export intracellular calcium using active transporters homologous to the sarco/endoplasmic reticulum Ca 2+ -ATPase (SERCA). Here we present three crystal structures of Ca 2+ -ATPase 1 from Listeria monocytogenes (LMCA1). Structures with BeF 3 - mimicking a phosphoenzyme state reveal a closed state, which is intermediate between the outward-open E2P and the proton-occluded E2-P* conformations known for SERCA. It suggests that LMCA1 in the E2P state is pre-organized for dephosphorylation upon Ca 2+ release, consistent with the rapid dephosphorylation observed in single-molecule studies. An arginine side-chain occupies the position equivalent to calcium binding site I in SERCA, leaving a single Ca 2+ binding site in LMCA1, corresponding to SERCA site II. Observing no putative transport pathways dedicated to protons, we infer a direct proton counter transport through the Ca 2+ exchange pathways. The LMCA1 structures provide insight into the evolutionary divergence and conserved features of this important class of ion transporters.
- Department of Molecular Biology and Genetics, Aarhus University, Denmark; The Danish Research Institute for Translational Neuroscience (DANDRITE), Nordic EMBL Partnership for Molecular Medicine, Denmark.
Organizational Affiliation: