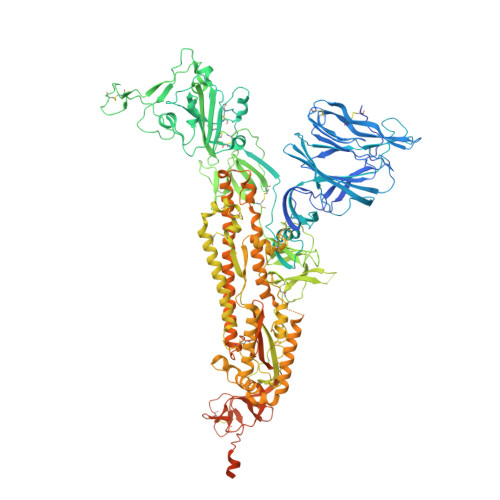
SARS-CoV-2 and bat RaTG13 spike glycoprotein structures inform on virus evolution and furin-cleavage effects.
Wrobel, A.G., Benton, D.J., Xu, P., Roustan, C., Martin, S.R., Rosenthal, P.B., Skehel, J.J., Gamblin, S.J.(2020) Nat Struct Mol Biol 27: 763-767
- PubMed: 32647346 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-020-0468-7
- Primary Citation Related Structures:
6ZGE, 6ZGF, 6ZGG, 6ZGH, 6ZGI - PubMed Abstract:
SARS-CoV-2 is thought to have emerged from bats, possibly via a secondary host. Here, we investigate the relationship of spike (S) glycoprotein from SARS-CoV-2 with the S protein of a closely related bat virus, RaTG13. We determined cryo-EM structures for RaTG13 S and for both furin-cleaved and uncleaved SARS-CoV-2 S; we compared these with recently reported structures for uncleaved SARS-CoV-2 S. We also biochemically characterized their relative stabilities and affinities for the SARS-CoV-2 receptor ACE2. Although the overall structures of human and bat virus S proteins are similar, there are key differences in their properties, including a more stable precleavage form of human S and about 1,000-fold tighter binding of SARS-CoV-2 to human receptor. These observations suggest that cleavage at the furin-cleavage site decreases the overall stability of SARS-CoV-2 S and facilitates the adoption of the open conformation that is required for S to bind to the ACE2 receptor.
- Structural Biology of Disease Processes Laboratory, Francis Crick Institute, London, UK. antoni.wrobel@crick.ac.uk.
Organizational Affiliation: