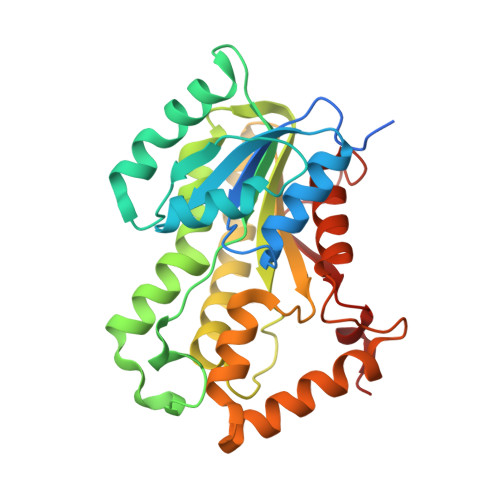
A Long Residence Time Enoyl-Reductase Inhibitor Explores an Extended Binding Region with Isoenzyme-Dependent Tautomer Adaptation and Differential Substrate-Binding Loop Closure.
Eltschkner, S., Kehrein, J., Le, T.A., Davoodi, S., Merget, B., Basak, S., Weinrich, J.D., Schiebel, J., Tonge, P.J., Engels, B., Sotriffer, C., Kisker, C.(2021) ACS Infect Dis 7: 746-758
- PubMed: 33710875 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsinfecdis.0c00437
- Primary Citation Related Structures:
6YUR, 6YUU - PubMed Abstract:
The enoyl-acyl carrier protein (ACP) reductase (ENR) is a key enzyme within the bacterial fatty-acid synthesis pathway. It has been demonstrated that small-molecule inhibitors carrying the diphenylether (DPE) scaffold bear a great potential for the development of highly specific and effective drugs against this enzyme class. Interestingly, different substitution patterns of the DPE scaffold have been shown to lead to varying effects on the kinetic and thermodynamic behavior toward ENRs from different organisms. Here, we investigated the effect of a 4'-pyridone substituent in the context of the slow tight-binding inhibitor SKTS1 on the inhibition of the Staphylococcus aureus enoyl-ACP-reductase saFabI and the closely related isoenzyme from Mycobacterium tuberculosis , InhA, and explored a new interaction site of DPE inhibitors within the substrate-binding pocket. Using high-resolution crystal structures of both complexes in combination with molecular dynamics (MD) simulations, kinetic measurements, and quantum mechanical (QM) calculations, we provide evidence that the 4'-pyridone substituent adopts different tautomeric forms when bound to the two ENRs. We furthermore elucidate the structural determinants leading to significant differences in the residence time of SKTS1 on both enzymes.
- Rudolf Virchow Center for Integrative and Translational Bioimaging, Institute for Structural Biology, University of Würzburg, 97080 Würzburg, Germany.
Organizational Affiliation: