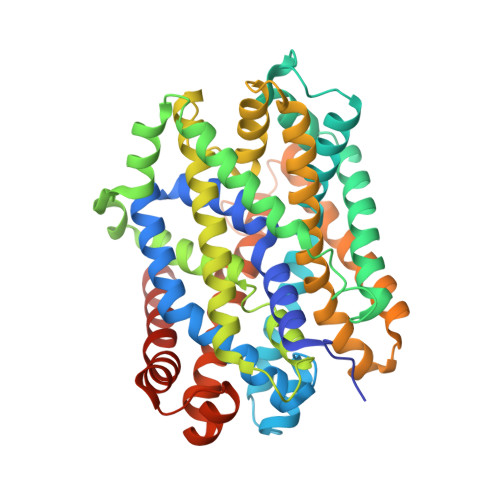
A non-helical region in transmembrane helix 6 of hydrophobic amino acid transporter MhsT mediates substrate recognition.
Focht, D., Neumann, C., Lyons, J., Eguskiza Bilbao, A., Blunck, R., Malinauskaite, L., Schwarz, I.O., Javitch, J.A., Quick, M., Nissen, P.(2021) EMBO J 40: e105164-e105164
- PubMed: 33155685 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2020105164
- Primary Citation Related Structures:
6YU2, 6YU3, 6YU4, 6YU5, 6YU6, 6YU7 - PubMed Abstract:
MhsT of Bacillus halodurans is a transporter of hydrophobic amino acids and a homologue of the eukaryotic SLC6 family of Na + -dependent symporters for amino acids, neurotransmitters, osmolytes, or creatine. The broad range of transported amino acids by MhsT prompted the investigation of the substrate recognition mechanism. Here, we report six new substrate-bound structures of MhsT, which, in conjunction with functional studies, reveal how the flexibility of a Gly-Met-Gly (GMG) motif in the unwound region of transmembrane segment 6 (TM6) is central for the recognition of substrates of different size by tailoring the binding site shape and volume. MhsT mutants, harboring substitutions within the unwound GMG loop and substrate binding pocket that mimick the binding sites of eukaryotic SLC6A18/B0AT3 and SLC6A19/B0AT1 transporters of neutral amino acids, exhibited impaired transport of aromatic amino acids that require a large binding site volume. Conservation of a general (G/A/C)ΦG motif among eukaryotic members of SLC6 family suggests a role for this loop in a common mechanism for substrate recognition and translocation by SLC6 transporters of broad substrate specificity.
- Department of Molecular Biology and Genetics, Danish Research Institute of Translational Neuroscience-DANDRITE, Nordic-EMBL Partnership for Molecular Medicine, Aarhus University, Aarhus C, Denmark.
Organizational Affiliation: