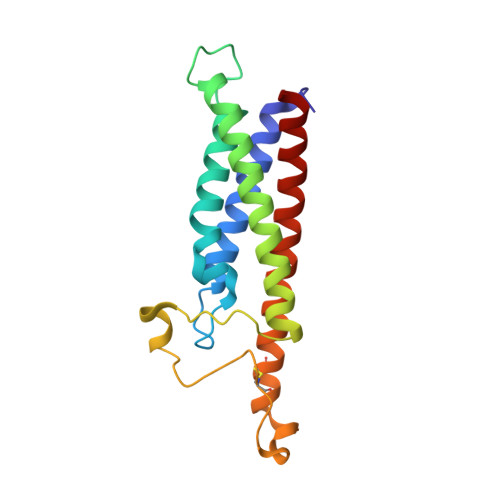
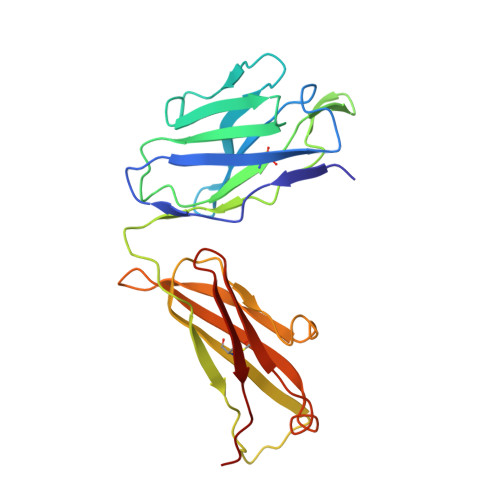
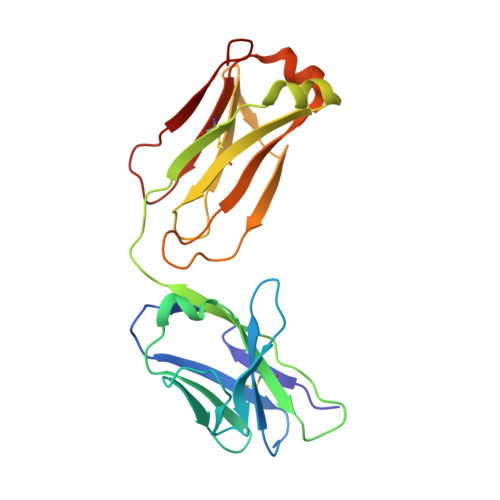
Binding mechanisms of therapeutic antibodies to human CD20.
Kumar, A., Planchais, C., Fronzes, R., Mouquet, H., Reyes, N.(2020) Science 369: 793-799
- PubMed: 32792392 Search on PubMed
- DOI: https://doi.org/10.1126/science.abb8008
- Primary Citation Related Structures:
6Y90, 6Y92, 6Y97, 6Y9A - PubMed Abstract:
Monoclonal antibodies (mAbs) targeting human antigen CD20 (cluster of differentiation 20) constitute important immunotherapies for the treatment of B cell malignancies and autoimmune diseases. Type I and II therapeutic mAbs differ in B cell binding properties and cytotoxic effects, reflecting differential interaction mechanisms with CD20. Here we present 3.7- to 4.7-angstrom cryo-electron microscopy structures of full-length CD20 in complexes with prototypical type I rituximab and ofatumumab and type II obinutuzumab. The structures and binding thermodynamics demonstrate that upon binding to CD20, type II mAbs form terminal complexes that preclude recruitment of additional mAbs and complement components, whereas type I complexes act as molecular seeds to increase mAb local concentration for efficient complement activation. Among type I mAbs, ofatumumab complexes display optimal geometry for complement recruitment. The uncovered mechanisms should aid rational design of next-generation immunotherapies targeting CD20.
- Membrane Protein Mechanisms Unit, Institut Pasteur, 75015 Paris, France.
Organizational Affiliation: