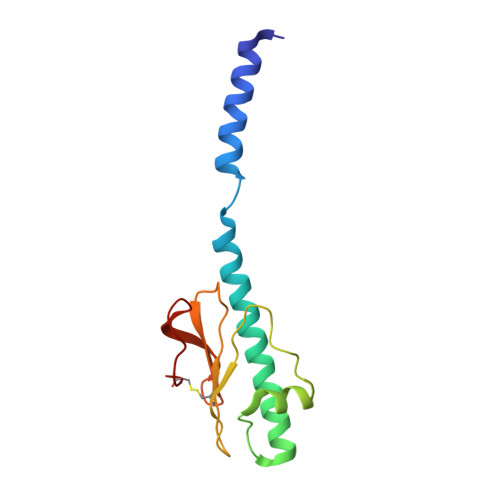
Cryo-electron microscopy reveals two distinct type IV pili assembled by the same bacterium.
Neuhaus, A., Selvaraj, M., Salzer, R., Langer, J.D., Kruse, K., Kirchner, L., Sanders, K., Daum, B., Averhoff, B., Gold, V.A.M.(2020) Nat Commun 11: 2231-2231
- PubMed: 32376942 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-15650-w
- Primary Citation Related Structures:
6XXD, 6XXE - PubMed Abstract:
Type IV pili are flexible filaments on the surface of bacteria, consisting of a helical assembly of pilin proteins. They are involved in bacterial motility (twitching), surface adhesion, biofilm formation and DNA uptake (natural transformation). Here, we use cryo-electron microscopy and mass spectrometry to show that the bacterium Thermus thermophilus produces two forms of type IV pilus ('wide' and 'narrow'), differing in structure and protein composition. Wide pili are composed of the major pilin PilA4, while narrow pili are composed of a so-far uncharacterized pilin which we name PilA5. Functional experiments indicate that PilA4 is required for natural transformation, while PilA5 is important for twitching motility.
- Living Systems Institute, University of Exeter, Stocker Road, Exeter, EX4 4QD, UK.
Organizational Affiliation: