Structure analysis of human SFPQ/NONO complex
Lee, M., Bond, C.S.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Splicing factor, proline- and glutamine-rich | 388 | Homo sapiens | Mutation(s): 0 Gene Names: SFPQ, PSF |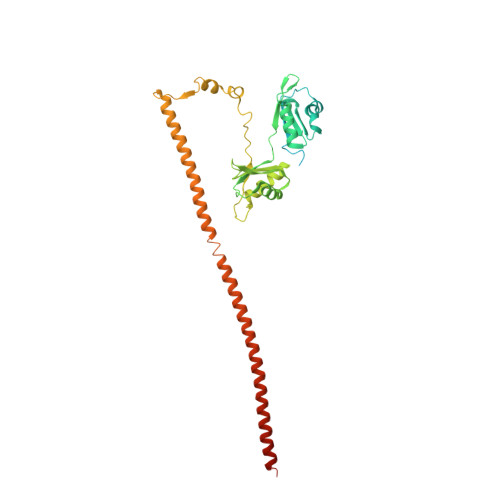  | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P23246 GTEx: ENSG00000116560 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P23246 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Non-POU domain-containing octamer-binding protein | 261 | Homo sapiens | Mutation(s): 0 Gene Names: NONO, NRB54 |  | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q15233 GTEx: ENSG00000147140 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q15233 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SO4 Download:Ideal Coordinates CCD File | E [auth D] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 66.63 | α = 90 |
| b = 112.94 | β = 90 |
| c = 204.62 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| MOSFLM | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Health and Medical Research Council (NHMRC, Australia) | Australia | 513935 |
| Australian Research Council (ARC) | Australia | DE150101243 |