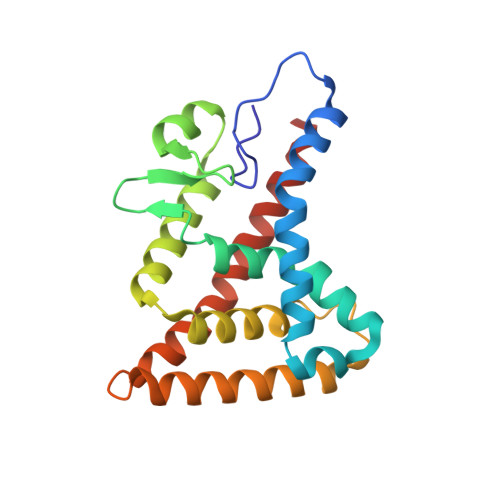
Structural basis for heme-dependent NCoR binding to the transcriptional repressor REV-ERB beta.
Mosure, S.A., Strutzenberg, T.S., Shang, J., Munoz-Tello, P., Solt, L.A., Griffin, P.R., Kojetin, D.J.(2021) Sci Adv 7
- PubMed: 33571111 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abc6479
- Primary Citation Related Structures:
6WMQ, 6WMS - PubMed Abstract:
Heme is the endogenous ligand for the constitutively repressive REV-ERB nuclear receptors, REV-ERBα (NR1D1) and REV-ERBβ (NR1D2), but how heme regulates REV-ERB activity remains unclear. Cellular studies indicate that heme is required for the REV-ERBs to bind the corepressor NCoR and repress transcription. However, fluorescence-based biochemical assays suggest that heme displaces NCoR; here, we show that this is due to a heme-dependent artifact. Using ITC and NMR spectroscopy, we show that heme binding remodels the thermodynamic interaction profile of NCoR receptor interaction domain (RID) binding to REV-ERBβ ligand-binding domain (LBD). We solved two crystal structures of REV-ERBβ LBD cobound to heme and NCoR peptides, revealing the heme-dependent NCoR binding mode. ITC and chemical cross-linking mass spectrometry reveals a 2:1 LBD:RID stoichiometry, consistent with cellular studies showing that NCoR-dependent repression of REV-ERB transcription occurs on dimeric DNA response elements. Our findings should facilitate renewed progress toward understanding heme-dependent REV-ERB activity.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, Jupiter, FL 33458, USA.
Organizational Affiliation: