Characterization of ArylalkylamineN-Acyltransferase fromTribolium castaneum: An Investigation into a Potential Next-Generation Insecticide Target.
O'Flynn, B.G., Lewandowski, E.M., Prins, K.C., Suarez, G., McCaskey, A.N., Rios-Guzman, N.M., Anderson, R.L., Shepherd, B.A., Gelis, I., Leahy, J.W., Chen, Y., Merkler, D.J.(2020) ACS Chem Biol 15: 513-523
- PubMed: 31967772 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.9b00973
- Primary Citation Related Structures:
6V3T - PubMed Abstract:
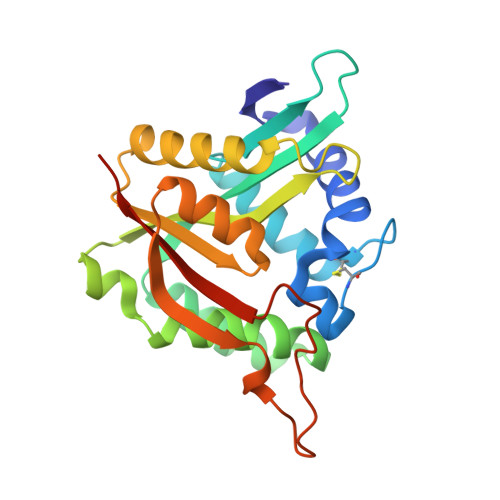
The growing issue of insecticide resistance has meant the identification of novel insecticide targets has never been more important. Arylalkylamine N -acyltransferases (AANATs) have been suggested as a potential new target. These promiscuous enzymes are involved in the N -acylation of biogenic amines to form N -acylamides. In insects, this process is a key step in melanism, hardening of the cuticle, removal of biogenic amines, and in the biosynthesis of fatty acid amides. The unique nature of each AANAT isoform characterized indicates each organism accommodates an assembly of discrete AANATs relatively exclusive to that organism. This implies a high potential for selectivity in insecticide design, while also maintaining polypharmacology. Presented here is a thorough kinetic and structural analysis of AANAT found in one of the most common secondary pests of all plant commodities in the world, Tribolium castaneum. The enzyme, named Tc AANAT0, catalyzes the formation of short-chain N -acylarylalkylamines, with short-chain acyl-CoAs (C2-C10), benzoyl-CoA, and succinyl-CoA functioning in the role of acyl donor. Recombinant Tc AANAT0 was expressed and purified from E. coli and was used to investigate the kinetic and chemical mechanism of catalysis. The kinetic mechanism is an ordered sequential mechanism with the acyl-CoA binding first. pH-rate profiles and site-directed mutagenesis studies identified amino acids critical to catalysis, providing insights about the chemical mechanism of Tc AANAT0. A crystal structure was obtained for Tc AANAT0 bound to acetyl-CoA, revealing valuable information about its active site. This combination of kinetic analysis and crystallography alongside mutagenesis and sequence analysis shines light on some approaches possible for targeting Tc AANAT0 and other AANATs for novel insecticide design.
- Department of Chemistry , University of South Florida , Tampa , Florida 33620 , United States.
Organizational Affiliation: