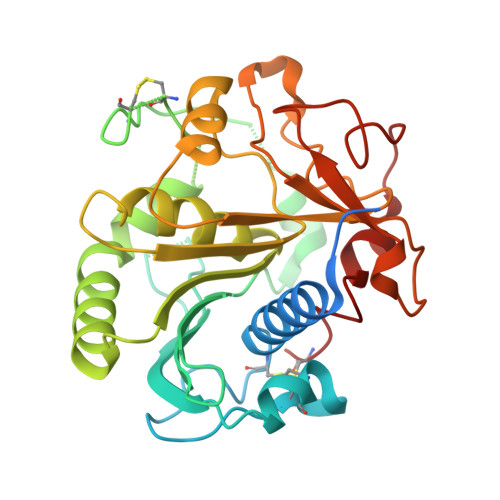
A Novel Fungal Lipase With Methanol Tolerance and Preference for Macaw Palm Oil.
Rade, L.L., da Silva, M.N.P., Vieira, P.S., Milan, N., de Souza, C.M., de Melo, R.R., Klein, B.C., Bonomi, A., de Castro, H.F., Murakami, M.T., Zanphorlin, L.M.(2020) Front Bioeng Biotechnol 8: 304-304
- PubMed: 32435636
- DOI: https://doi.org/10.3389/fbioe.2020.00304
- Primary Citation Related Structures:
6UNV - PubMed Abstract:
Macaw palm is a highly oil-producing plant, which presents high contents of free fatty acids, being a promising feedstock for biofuel production. The current chemical routes are costly and complex, involving highly harsh industrial conditions. Enzymatic processing is a potential alternative; however, it is hampered by the scarce knowledge on biocatalysts adapted to this acidic feedstock. This work describes a novel lipase isolated from the thermophilic fungus Rasamsonia emersonii ( Re Lip), which tolerates extreme conditions such as the presence of methanol, high temperatures, and acidic medium. Among the tested feedstocks, the enzyme showed the highest preference for macaw palm oil, producing a hydrolyzate with a final free fatty acid content of 92%. Crystallographic studies revealed a closed conformation of the helical amphipathic lid that typically undergoes conformational changes in a mechanism of interfacial activation. Such conformation of the lid is stabilized by a salt bridge, not observed in other structurally characterized homologs, which is likely involved in the tolerance to organic solvents. Moreover, the lack of conservation of the aromatic cluster IxxWxxxxxF in the lid of Re Lip with the natural mutation of the phenylalanine by an alanine might be correlated with the preference of short acyl chains, although preserving catalytic activity on insoluble substrates. In addition, the presence of five acidic amino acids in the lid of Re Lip, a rare property reported in other lipases, may have contributed to its ability to tolerate and be effective in acidic environments. Therefore, our work describes a new fungal biocatalyst capable of efficiently hydrolyzing macaw oil, an attractive feedstock for the production of "drop-in" biofuels, with high desirable feature for industrial conditions such as thermal and methanol tolerance, and optimum acidic pH. Moreover, the crystallographic structure was elucidated, providing a structural basis for the enzyme substrate preference and tolerance to organic solvents.
- Brazilian Biorenewables National Laboratory, Brazilian Center for Research in Energy and Materials, Campinas, Brazil.
Organizational Affiliation: