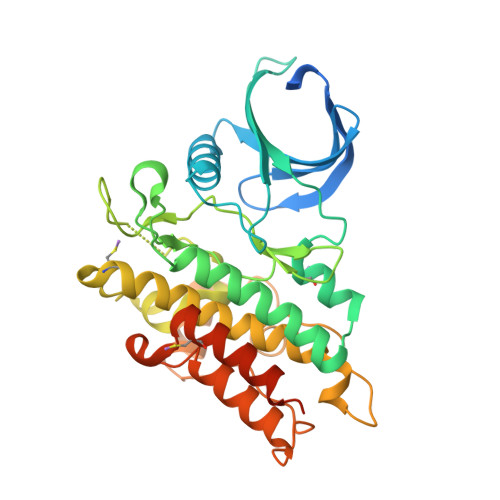
Structural basis for ALK2/BMPR2 receptor complex signaling through kinase domain oligomerization.
Agnew, C., Ayaz, P., Kashima, R., Loving, H.S., Ghatpande, P., Kung, J.E., Underbakke, E.S., Shan, Y., Shaw, D.E., Hata, A., Jura, N.(2021) Nat Commun 12: 4950-4950
- PubMed: 34400635 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-25248-5
- Primary Citation Related Structures:
6UNP, 6UNQ, 6UNR, 6UNS - PubMed Abstract:
Upon ligand binding, bone morphogenetic protein (BMP) receptors form active tetrameric complexes, comprised of two type I and two type II receptors, which then transmit signals to SMAD proteins. The link between receptor tetramerization and the mechanism of kinase activation, however, has not been elucidated. Here, using hydrogen deuterium exchange mass spectrometry (HDX-MS), small angle X-ray scattering (SAXS) and molecular dynamics (MD) simulations, combined with analysis of SMAD signaling, we show that the kinase domain of the type I receptor ALK2 and type II receptor BMPR2 form a heterodimeric complex via their C-terminal lobes. Formation of this dimer is essential for ligand-induced receptor signaling and is targeted by mutations in BMPR2 in patients with pulmonary arterial hypertension (PAH). We further show that the type I/type II kinase domain heterodimer serves as the scaffold for assembly of the active tetrameric receptor complexes to enable phosphorylation of the GS domain and activation of SMADs.
- Cardiovascular Research Institute, University of California San Francisco, San Francisco, CA, USA.
Organizational Affiliation: