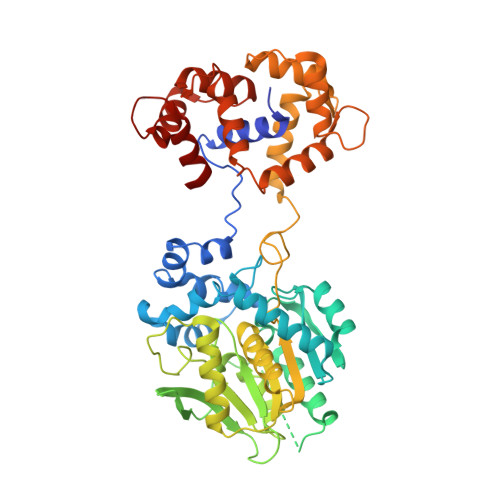
The HaloTag as a general scaffold for far-red tunable chemigenetic indicators.
Deo, C., Abdelfattah, A.S., Bhargava, H.K., Berro, A.J., Falco, N., Farrants, H., Moeyaert, B., Chupanova, M., Lavis, L.D., Schreiter, E.R.(2021) Nat Chem Biol 17: 718-723
- PubMed: 33795886 Search on PubMed
- DOI: https://doi.org/10.1038/s41589-021-00775-w
- Primary Citation Related Structures:
6U2M, 6U32 - PubMed Abstract:
Functional imaging using fluorescent indicators has revolutionized biology, but additional sensor scaffolds are needed to access properties such as bright, far-red emission. Here, we introduce a new platform for 'chemigenetic' fluorescent indicators, utilizing the self-labeling HaloTag protein conjugated to environmentally sensitive synthetic fluorophores. We solve a crystal structure of HaloTag bound to a rhodamine dye ligand to guide engineering efforts to modulate the dye environment. We show that fusion of HaloTag with protein sensor domains that undergo conformational changes near the bound dye results in large and rapid changes in fluorescence output. This generalizable approach affords bright, far-red calcium and voltage sensors with highly tunable photophysical and chemical properties, which can reliably detect single action potentials in cultured neurons.
- Janelia Research Campus, Howard Hughes Medical Institute, Ashburn, VA, USA.
Organizational Affiliation: