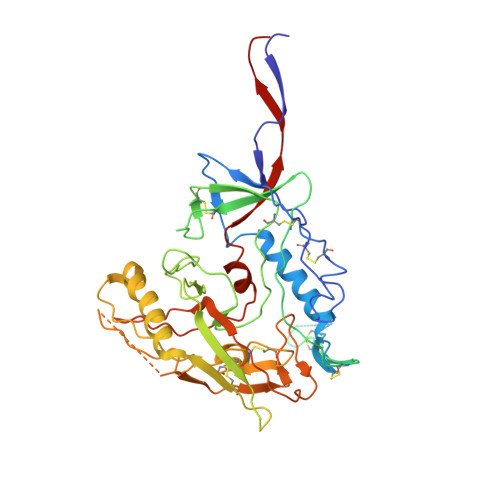
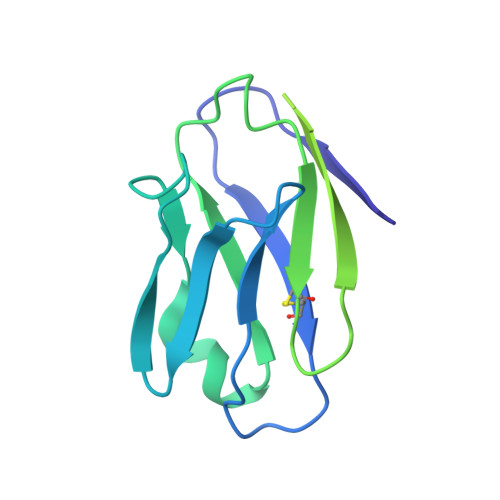
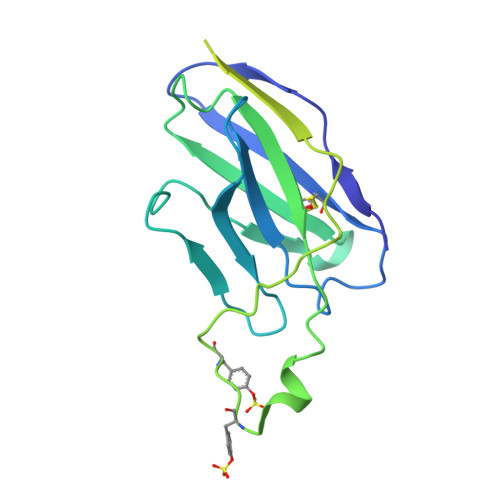
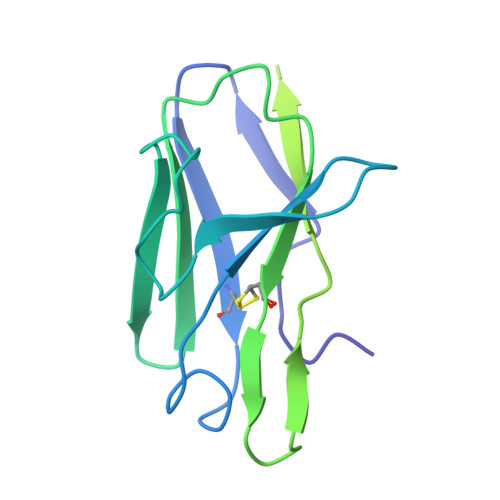
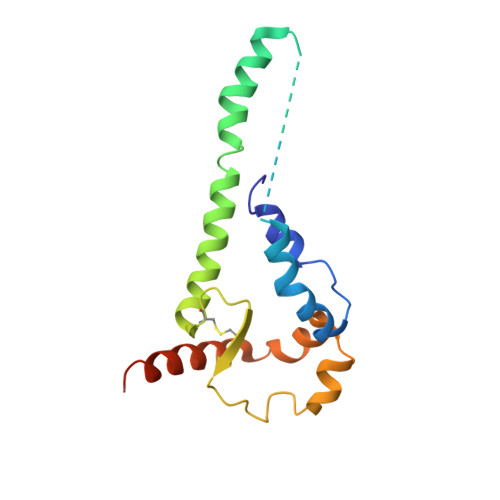
Asymmetric opening of HIV-1 Env bound to CD4 and a coreceptor-mimicking antibody.
Yang, Z., Wang, H., Liu, A.Z., Gristick, H.B., Bjorkman, P.J.(2019) Nat Struct Mol Biol 26: 1167-1175
- PubMed: 31792452 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-019-0344-5
- Primary Citation Related Structures:
6U0L, 6U0N - PubMed Abstract:
The human immunodeficiency virus (HIV-1) envelope (Env) glycoprotein, a (gp120-gp41) 3 trimer, mediates fusion of viral and host cell membranes after gp120 binding to host receptor CD4. Receptor binding triggers conformational changes allowing coreceptor (CCR5) recognition through CCR5's tyrosine-sulfated amino (N) terminus, release of the gp41 fusion peptide and fusion. We present 3.3 Å and 3.5 Å cryo-EM structures of E51, a tyrosine-sulfated coreceptor-mimicking antibody, complexed with a CD4-bound open HIV-1 native-like Env trimer. Two classes of asymmetric Env interact with E51, revealing tyrosine-sulfated interactions with gp120 mimicking CCR5 interactions, and two conformations of gp120-gp41 protomers (A and B protomers in AAB and ABB trimers) that differ in their degree of CD4-induced trimer opening and induction of changes to the fusion peptide. By integrating the new structural information with previous closed and open envelope trimer structures, we modeled the order of conformational changes on the path to coreceptor binding site exposure and subsequent viral-host cell membrane fusion.
- Division of Biology and Biological Engineering, California Institute of Technology, Pasadena CA, USA.
Organizational Affiliation: