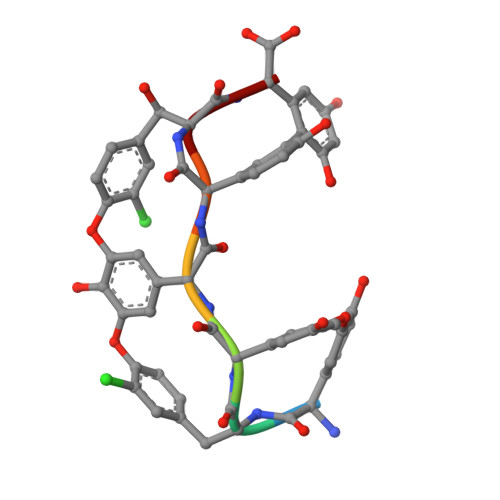
Enantiomeric Separation and Molecular Modelling of Bioactive 4-Aryl-3,4-dihydropyrimidin-2(1H)-one Ester Derivatives on Teicoplanin-Based Chiral Stationary Phase
Bolognino, I., Carrieri, A., Purgatorio, R., Catto, M., Caliandro, R., Carrozzini, B., Belviso, B.D., Majellaro, M., Sotelo, E., Cellamare, S., Altomare, C.D.(2022) Separations