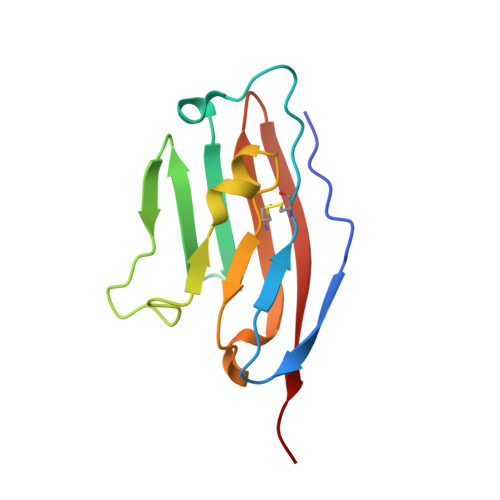
Human and mouse PD-L1: similar molecular structure, but different druggability profiles.
Magiera-Mularz, K., Kocik, J., Musielak, B., Plewka, J., Sala, D., Machula, M., Grudnik, P., Hajduk, M., Czepiel, M., Siedlar, M., Holak, T.A., Skalniak, L.(2021) iScience 24: 101960-101960
- PubMed: 33437940 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.isci.2020.101960
- Primary Citation Related Structures:
6SRU - PubMed Abstract:
In the development of PD-L1-blocking therapeutics, it is essential to transfer initial in vitro findings into proper in vivo animal models. Classical immunocompetent mice are attractive due to high accessibility and low experimental costs. However, it is unknown whether inter-species differences in PD-L1 sequence and structure would allow for human-mouse cross applications. Here, we disclose the first structure of the mouse ( m ) PD-L1 and analyze its similarity to the human ( h ) PD-L1. We show that m PD-L1 interacts with h PD-1 and provides a negative signal toward activated Jurkat T cells. We also show major differences in druggability between the h PD-L1 and m PD-L1 using therapeutic antibodies, a macrocyclic peptide, and small molecules. Our study indicates that while the amino acid sequence is well conserved between the h PD-L1 and m PD-L1 and overall structures are almost identical, crucial differences determine the interaction with anti-PD-L1 agents, that cannot be easily predicted in silico .
- Department of Organic Chemistry, Faculty of Chemistry, Jagiellonian University, Gronostajowa 2, 30-387 Krakow, Poland.
Organizational Affiliation: