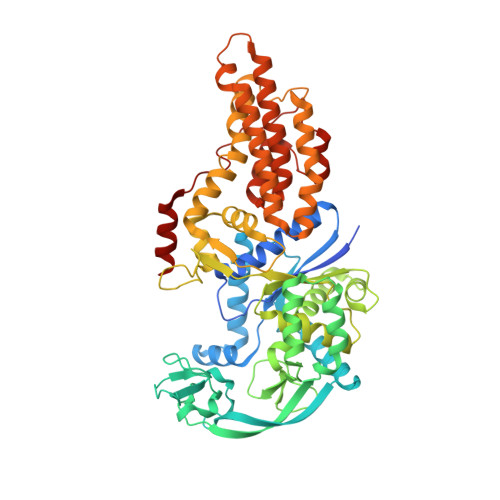
Use of beta3-methionine as an amino acid substrate of Escherichia coli methionyl-tRNA synthetase.
Nigro, G., Bourcier, S., Lazennec-Schurdevin, C., Schmitt, E., Marliere, P., Mechulam, Y.(2020) J Struct Biol 209: 107435-107435
- PubMed: 31862305 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2019.107435
- Primary Citation Related Structures:
6SPN, 6SPO, 6SPP, 6SPQ, 6SPR - PubMed Abstract:
Polypeptides containing β-amino acids are attractive tools for the design of novel proteins having unique properties of medical or industrial interest. Incorporation of β-amino acids in vivo requires the development of efficient aminoacyl-tRNA synthetases specific of these non-canonical amino acids. Here, we have performed a detailed structural and biochemical study of the recognition and use of β 3 -Met by Escherichia coli methionyl-tRNA synthetase (MetRS). We show that MetRS binds β 3 -Met with a 24-fold lower affinity but catalyzes the esterification of the non-canonical amino acid onto tRNA with a rate lowered by three orders of magnitude. Accurate measurements of the catalytic parameters required careful consideration of the presence of contaminating α-Met in β 3 -Met commercial samples. The 1.45 Å crystal structure of the MetRS: β 3 -Met complex shows that β 3 -Met binds the enzyme essentially like α-Met, but the carboxylate moiety is mobile and not adequately positioned to react with ATP for aminoacyl adenylate formation. This study provides structural and biochemical bases for engineering MetRS with improved β 3 -Met aminoacylation capabilities.
- Laboratoire de Biochimie, BIOC, Ecole polytechnique, CNRS, Institut Polytechnique de Paris, 91128 Palaiseau Cedex, France.
Organizational Affiliation: