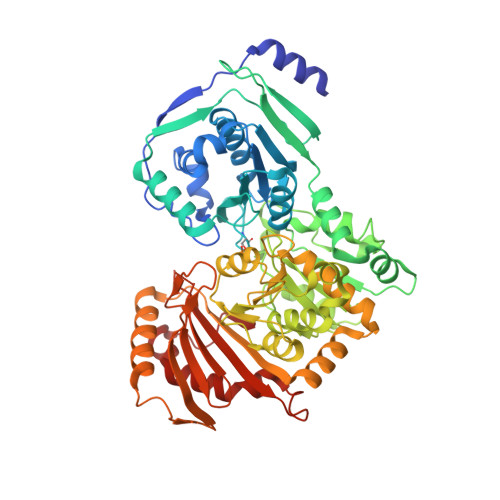
Structural basis for substrate and product recognition in human phosphoglucomutase-1 (PGM1) isoform 2, a member of the alpha-D-phosphohexomutase superfamily.
Backe, P.H., Laerdahl, J.K., Kittelsen, L.S., Dalhus, B., Morkrid, L., Bjoras, M.(2020) Sci Rep 10: 5656-5656
- PubMed: 32221390
- DOI: https://doi.org/10.1038/s41598-020-62548-0
- Primary Citation Related Structures:
6SNO, 6SNP, 6SNQ - PubMed Abstract:
Human phosphoglucomutase 1 (PGM1) is an evolutionary conserved enzyme that belongs to the ubiquitous and ancient α-D-phosphohexomutases, a large enzyme superfamily with members in all three domains of life. PGM1 catalyzes the bi-directional interconversion between α-D-glucose 1-phosphate (G1P) and α-D-glucose 6-phosphate (G6P), a reaction that is essential for normal carbohydrate metabolism and also important in the cytoplasmic biosynthesis of nucleotide sugars needed for glycan biosynthesis. Clinical studies have shown that mutations in the PGM1 gene may cause PGM1 deficiency, an inborn error of metabolism previously classified as a glycogen storage disease, and PGM1 deficiency was recently also shown to be a congenital disorder of glycosylation. Here we present three crystal structures of the isoform 2 variant of PGM1, both as a free enzyme and in complex with its substrate and product. The structures show the longer N-terminal of this PGM1 variant, and the ligand complex structures reveal for the first time the detailed structural basis for both G1P substrate and G6P product recognition by human PGM1. We also show that PGM1 and the paralogous gene PGM5 are the results of a gene duplication event in a common ancestor of jawed vertebrates, and, importantly, that both PGM1 isoforms are conserved and of functional significance in all vertebrates. Our finding that PGM1 encodes two equally conserved and functionally important isoforms in the human organism should be taken into account in the evaluation of disease-related missense mutations in patients in the future.
- Department of Microbiology, Oslo University Hospital, Oslo, Norway. paul.hoff.backe@rr-research.no.
Organizational Affiliation: