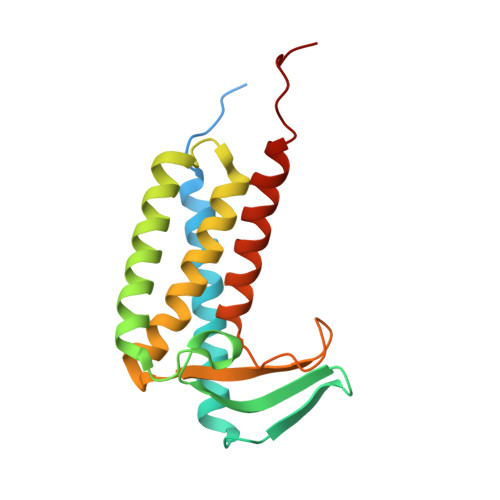
Structures of lipoprotein signal peptidase II from Staphylococcus aureus complexed with antibiotics globomycin and myxovirescin.
Olatunji, S., Yu, X., Bailey, J., Huang, C.Y., Zapotoczna, M., Bowen, K., Remskar, M., Muller, R., Scanlan, E.M., Geoghegan, J.A., Olieric, V., Caffrey, M.(2020) Nat Commun 11: 140-140
- PubMed: 31919415 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-13724-y
- Primary Citation Related Structures:
6RYO, 6RYP - PubMed Abstract:
Antimicrobial resistance is a major global threat that calls for new antibiotics. Globomycin and myxovirescin are two natural antibiotics that target the lipoprotein-processing enzyme, LspA, thereby compromising the integrity of the bacterial cell envelope. As part of a project aimed at understanding their mechanism of action and for drug development, we provide high-resolution crystal structures of the enzyme from the human pathogen methicillin-resistant Staphylococcus aureus (MRSA) complexed with globomycin and with myxovirescin. Our results reveal an instance of convergent evolution. The two antibiotics possess different molecular structures. Yet, they appear to inhibit identically as non-cleavable tetrahedral intermediate analogs. Remarkably, the two antibiotics superpose along nineteen contiguous atoms that interact similarly with LspA. This 19-atom motif recapitulates a part of the substrate lipoprotein in its proposed binding mode. Incorporating this motif into a scaffold with suitable pharmacokinetic properties should enable the development of effective antibiotics with built-in resistance hardiness.
- Membrane Structural and Functional Biology Group, School of Medicine and School of Biochemistry and Immunology, Trinity College Dublin, Dublin, D02 R590, Ireland.
Organizational Affiliation: