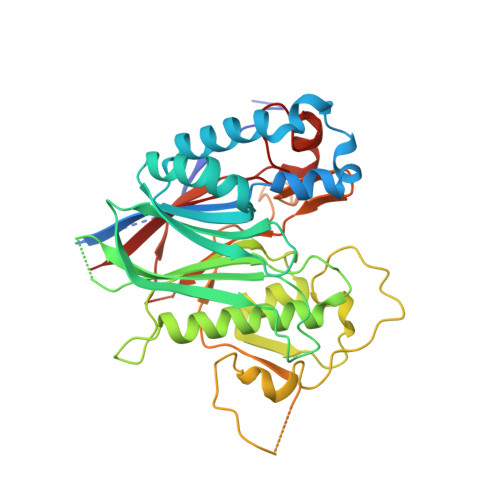
ANGEL2 is a member of the CCR4 family of deadenylases with 2',3'-cyclic phosphatase activity.
Pinto, P.H., Kroupova, A., Schleiffer, A., Mechtler, K., Jinek, M., Weitzer, S., Martinez, J.(2020) Science 369: 524-530
- PubMed: 32732418 Search on PubMed
- DOI: https://doi.org/10.1126/science.aba9763
- Primary Citation Related Structures:
6RVZ, 6RW0 - PubMed Abstract:
RNA molecules are frequently modified with a terminal 2',3'-cyclic phosphate group as a result of endonuclease cleavage, exonuclease trimming, or de novo synthesis. During pre-transfer RNA (tRNA) and unconventional messenger RNA (mRNA) splicing, 2',3'-cyclic phosphates are substrates of the tRNA ligase complex, and their removal is critical for recycling of tRNAs upon ribosome stalling. We identified the predicted deadenylase angel homolog 2 (ANGEL2) as a human phosphatase that converts 2',3'-cyclic phosphates into 2',3'-OH nucleotides. We analyzed ANGEL2's substrate preference, structure, and reaction mechanism. Perturbing ANGEL2 expression affected the efficiency of pre-tRNA processing, X-box-binding protein 1 ( XBP1 ) mRNA splicing during the unfolded protein response, and tRNA nucleotidyltransferase 1 (TRNT1)-mediated CCA addition onto tRNAs. Our results indicate that ANGEL2 is involved in RNA pathways that rely on the ligation or hydrolysis of 2',3'-cyclic phosphates.
- Max Perutz Labs, Medical University of Vienna, Vienna BioCenter, 1030 Vienna, Austria.
Organizational Affiliation: