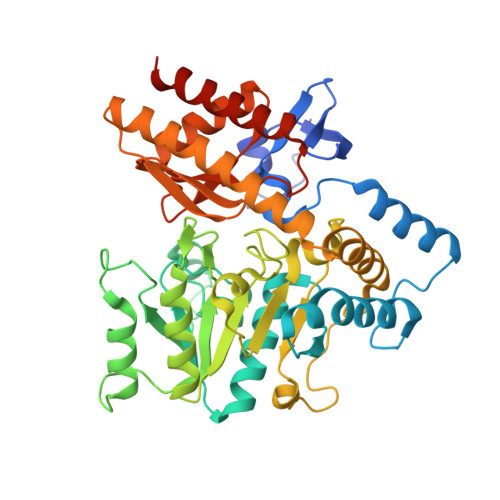
The crystal structure of the tetrameric DABA-aminotransferase EctB, a rate-limiting enzyme in the ectoine biosynthesis pathway.
Hillier, H.T., Altermark, B., Leiros, I.(2020) FEBS J 287: 4641-4658
- PubMed: 32112674 Search on PubMed
- DOI: https://doi.org/10.1111/febs.15265
- Primary Citation Related Structures:
6RL5 - PubMed Abstract:
l-2,4-diaminobutyric acid (DABA) aminotransferases can catalyze the formation of amines at the distal ω-position of substrates, and is the intial and rate-limiting enzyme in the biosynthesis pathway of the cytoprotecting molecule (S)-2-methyl-1,4,5,6-tetrahydro-4-pyrimidine carboxylic acid (ectoine). Although there is an industrial interest in the biosynthesis of ectoine, the DABA aminotransferases remain poorly characterized. Herein, we present the crystal structure of EctB (2.45 Å), a DABA aminotransferase from Chromohalobacter salexigens DSM 3043, a well-studied organism with respect to osmoadaptation by ectoine biosynthesis. We investigate the enzyme's oligomeric state to show that EctB from C. salexigens is a tetramer of two functional dimers, and suggest conserved recognition sites for dimerization that also includes the characteristic gating loop that helps shape the active site of the neighboring monomer. Although ω-transaminases are known to have two binding pockets to accommodate for their dual substrate specificity, we herein provide the first description of two binding pockets in the active site that may account for the catalytic character of DABA aminotransferases. Furthermore, our biochemical data reveal that the EctB enzyme from C. salexigens is a thermostable, halotolerant enzyme with a broad pH tolerance which may be linked to its tetrameric state. Put together, this study creates a solid foundation for a deeper structural understanding of DABA aminotransferases and opening up for future downstream studies of EctB's catalytic character and its redesign as a better catalyst for ectoine biosynthesis. In summary, we believe that the EctB enzyme from C. salexigens can serve as a benchmark enzyme for characterization of DABA aminotransferases. DATABASE: Structural data are available in PDB database under the accession number 6RL5.
- The Norwegian Structural Biology Centre (NorStruct), Department of Chemistry, Faculty of Science and Technology, UiT the Arctic University of Norway, Tromsø, Norway.
Organizational Affiliation: