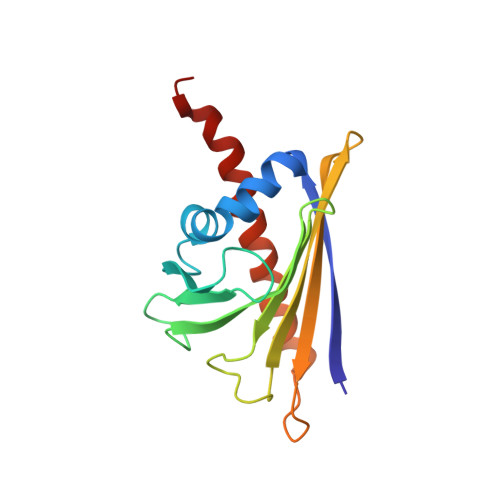
Identification of a natural ligand of the hazel allergen Cor a 1.
Jacob, T., von Loetzen, C.S., Reuter, A., Lacher, U., Schiller, D., Schobert, R., Mahler, V., Vieths, S., Rosch, P., Schweimer, K., Wohrl, B.M.(2019) Sci Rep 9: 8714-8714
- PubMed: 31213622 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-44999-2
- Primary Citation Related Structures:
6GQ9, 6R3C - PubMed Abstract:
Hazelnut is one of the most frequent causes of food allergy. The major hazel allergen in Northern Europe is Cor a 1, which is homologous to the major birch pollen allergen Bet v 1. Both allergens belong to the pathogenesis related class PR-10. We determined the solution structure of Cor a 1.0401 from hazelnut and identified a natural ligand of the protein. The structure reveals the protein fold characteristic for PR-10 family members, which consists of a seven-stranded antiparallel β-sheet, two short α-helices arranged in V-shape and a long C-terminal α-helix encompassing a hydrophobic pocket. However, despite the structural similarities between Cor a 1 and Bet v 1, they bind different ligands. We have shown previously that Bet v 1 binds to quercetin-3-O-sophoroside. Here, we isolated Cor a 1 from hazel pollen and identified the bound ligand, quercetin-3-O-(2"-O-β-D-glucopyranosyl)-β-D-galactopyranoside, by mass spectrometry and nuclear magnetic resonance spectroscopy (NMR). NMR experiments were performed to confirm binding. Remarkably, although it has been shown that PR-10 allergens show promiscuous binding behaviour in vitro, we can demonstrate that Cor a 1.0401 and Bet v 1.0101 exhibit highly selective binding for their specific ligand but not for the respective ligand of the other allergen.
- Universität Bayreuth, Lehrstuhl Biopolymere, Bayreuth, Germany.
Organizational Affiliation: