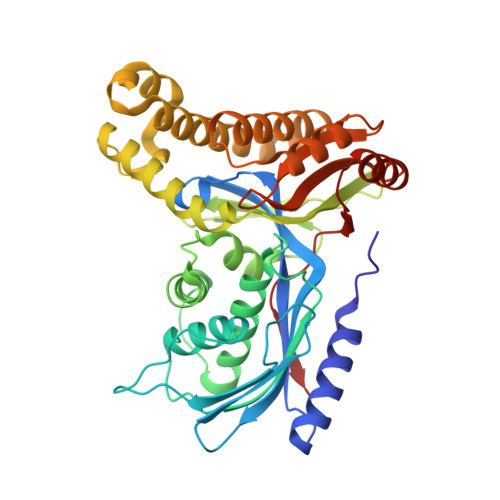
Structure of human galactokinase 1 bound with 4-{[2-(Methylsulfonyl)-1H-imidazol-1-yl]methyl}-1,3-thiazole
Mackinnon, S.R., Bezerra, G.A., Zhang, M., Foster, W., Krojer, T., Brandao-Neto, J., Douangamath, A., Arrowsmith, C., Edwards, A., Bountra, C., Brennan, P., Lai, K., Yue, W.W.To be published.