Odorant-binding site in visual opsin
Morizumi, T., Kuroi, K., Eger, B.T., Ou, W.L., Van Eps, N., Tsukamoto, H., Furutani, Y., Ernst, O.P.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Rhodopsin | 348 | Bos taurus | Mutation(s): 0 Membrane Entity: Yes | 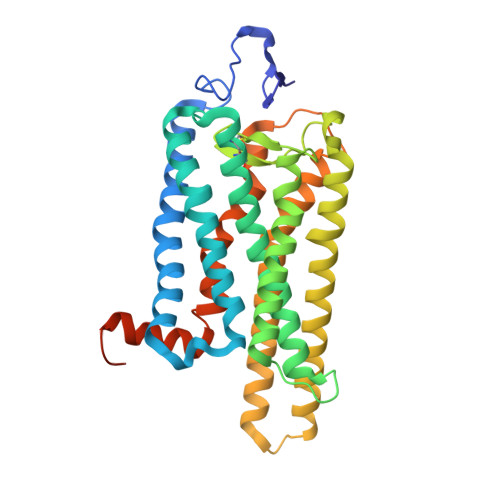 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P02699 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | |||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| ILENLKDVGLF G alpha peptide CT2 | 11 | Bos taurus | Mutation(s): 0 |  | |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| beta-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | C | 4 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G31886NL GlyCosmos: G31886NL GlyGen: G31886NL | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| BOG Download:Ideal Coordinates CCD File | E [auth A] | octyl beta-D-glucopyranoside C14 H28 O6 HEGSGKPQLMEBJL-RKQHYHRCSA-N |  | ||
| PLM Download:Ideal Coordinates CCD File | F [auth A] | PALMITIC ACID C16 H32 O2 IPCSVZSSVZVIGE-UHFFFAOYSA-N |  | ||
| ODM (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | D [auth A] | (3R)-3,7-dimethyloct-6-en-1-ol C10 H20 O QMVPMAAFGQKVCJ-SNVBAGLBSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 243.412 | α = 90 |
| b = 243.412 | β = 90 |
| c = 108.44 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| autoPROC | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Canadian Institutes of Health Research (CIHR) | Canada | Canada Research Excellence Chair |