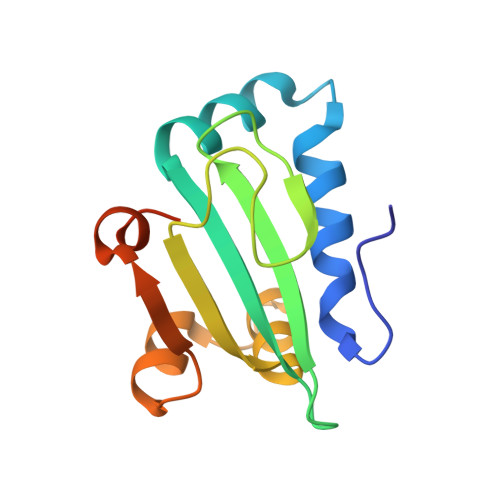
The structure of unliganded sterol carrier protein 2 from Yarrowia lipolytica unveils a mechanism for binding site occlusion.
Gianotti, A.R., Klinke, S., Ermacora, M.R.(2020) J Struct Biol 213: 107675-107675
- PubMed: 33278583 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2020.107675
- Primary Citation Related Structures:
6OVP - PubMed Abstract:
Isolated or as a part of multidomain proteins, Sterol Carrier Protein 2 (SCP2) exhibits high affinity and broad specificity for different lipidic and hydrophobic compounds. A wealth of structural information on SCP2 domains in all forms of life is currently available; however, many aspects of its ligand binding activity are poorly understood. ylSCP2 is a well-characterized single domain SCP2 from the yeast Yarrowia lipolytica. Herein, we report the X-ray structure of unliganded ylSCP2 refined to 2.0 Å resolution. Comparison with the previously solved liganded ylSCP2 structure unveiled a novel mechanism for binding site occlusion. The liganded ylSCP2 binding site is a large cavity with a volume of more than 800 Å 3 . In unliganded ylSCP2 the binding site is reduced to about 140 Å 3 . The obliteration is caused by a swing movement of the C-terminal α helix 5 and a subtle compaction of helices 2-4. Previous pairwise comparisons were between homologous SCP2 domains with a uncertain binding status. The reported unliganded ylSCP2 structure allows for the first time a fully controlled comparative analysis of the conformational effects of ligand occupation dispelling several doubts regarding the architecture of SCP2 binding site.
- Departamento de Ciencia y Tecnología, Universidad Nacional de Quilmes, Argentina; Grupo de Biología Estructural y Biotecnología, IMBICE, CONICET, Universidad Nacional de Quilmes, Argentina.
Organizational Affiliation: