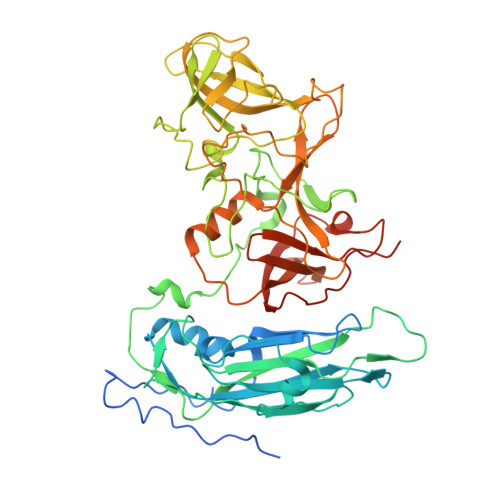
High-resolution cryo-EM structures of outbreak strain human norovirus shells reveal size variations.
Jung, J., Grant, T., Thomas, D.R., Diehnelt, C.W., Grigorieff, N., Joshua-Tor, L.(2019) Proc Natl Acad Sci U S A 116: 12828-12832
- PubMed: 31182604 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1903562116
- Primary Citation Related Structures:
6OTF, 6OU9, 6OUC, 6OUT, 6OUU - PubMed Abstract:
Noroviruses are a leading cause of foodborne illnesses worldwide. Although GII.4 strains have been responsible for most norovirus outbreaks, the assembled virus shell structures have been available in detail for only a single strain (GI.1). We present high-resolution (2.6- to 4.1-Å) cryoelectron microscopy (cryo-EM) structures of GII.4, GII.2, GI.7, and GI.1 human norovirus outbreak strain virus-like particles (VLPs). Although norovirus VLPs have been thought to exist in a single-sized assembly, our structures reveal polymorphism between and within genogroups, with small, medium, and large particle sizes observed. Using asymmetric reconstruction, we were able to resolve a Zn 2+ metal ion adjacent to the coreceptor binding site, which affected the structural stability of the shell. Our structures serve as valuable templates for facilitating vaccine formulations.
- W. M. Keck Structural Biology Laboratory, Cold Spring Harbor Laboratory, Cold Spring Harbor, NY 11724.
Organizational Affiliation: