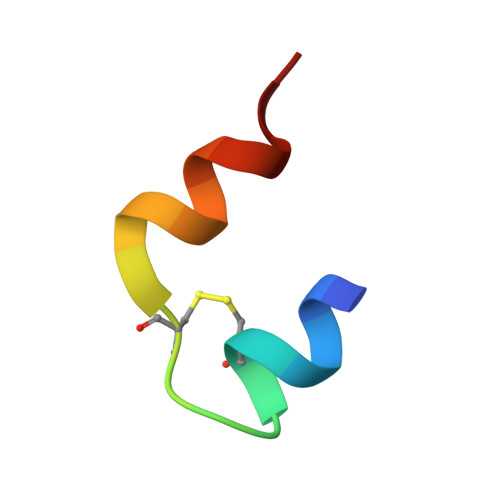
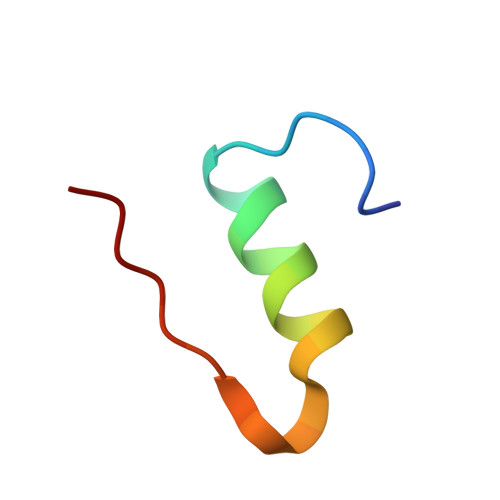
Non-merohedral twinning: from minerals to proteins.
Sevvana, M., Ruf, M., Uson, I., Sheldrick, G.M., Herbst-Irmer, R.(2019) Acta Crystallogr D Struct Biol 75: 1040-1050
- PubMed: 31793898 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798319010179
- Primary Citation Related Structures:
6OQZ, 6OR0 - PubMed Abstract:
In contrast to twinning by merohedry, the reciprocal lattices of the different domains of non-merohedral twins do not overlap exactly. This leads to three kinds of reflections: reflections with no overlap, reflections with an exact overlap and reflections with a partial overlap of a reflection from a second domain. This complicates the unit-cell determination, indexing, data integration and scaling of X-ray diffraction data. However, with hindsight it is possible to detwin the data because there are reflections that are not affected by the twinning. In this article, the successful solution and refinement of one mineral, one organometallic and two protein non-merohedral twins using a common strategy are described. The unit-cell constants and the orientation matrices were determined by the program CELL_NOW. The data were then integrated with SAINT. TWINABS was used for scaling, empirical absorption corrections and the generation of two different data files, one with detwinned data for structure solution and refinement and a second one for (usually more accurate) structure refinement against total integrated intensities. The structures were solved by experimental phasing using SHELXT for the first two structures and SHELXC/D/E for the two protein structures; all models were refined with SHELXL.
- Department of Biological Sciences, Purdue University, West Lafayette, IN 47907, USA.
Organizational Affiliation: