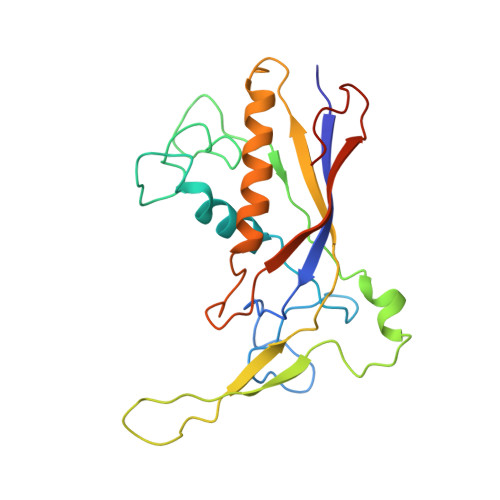
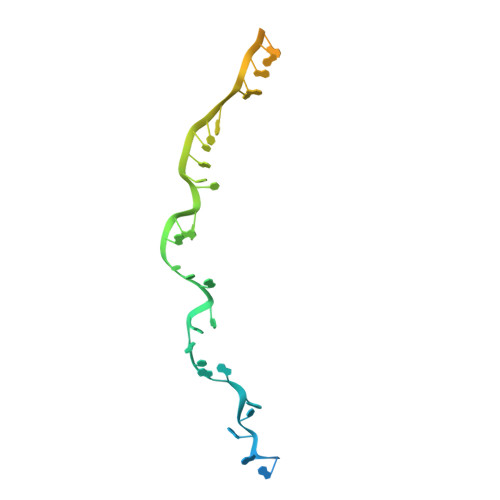
Coupling of ssRNA cleavage with DNase activity in type III-A CRISPR-Csm revealed by cryo-EM and biochemistry.
Guo, M., Zhang, K., Zhu, Y., Pintilie, G.D., Guan, X., Li, S., Schmid, M.F., Ma, Z., Chiu, W., Huang, Z.(2019) Cell Res 29: 305-312
- PubMed: 30814678 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-019-0151-x
- Primary Citation Related Structures:
6NUD, 6NUE - PubMed Abstract:
The type III CRISPR-Cas (clustered regularly interspaced short palindromic repeats-CRISPR-associated genes) systems are bacterially encoded adaptive immune systems for defense against invading nucleic acids. They accomplish this task through the coordinated cleavage of invading substrates of single-stranded RNA and DNA (ssDNA and ssRNA) by the Csm (type III-A) or Cmr (type III-B) effector complexes. The ssRNA is complementarily bound to the CRISPR RNA (crRNA). However, the structural basis for the DNase and RNase activation of the Csm nucleoprotein complex is largely unknown. Here we report cryo-EM structures of the Csm-crRNA complex, with or without target ssRNA, at near-atomic resolution. Our cryo-EM maps allow us to build atomic models of the key macromolecular components, including Cas10, Csm2, Csm3, Csm4, crRNA and the invading ssRNA. Our structure resolves unambiguously the stoichiometry and tertiary structures of the Csm protein complex and the interactions between protein components and the crRNA/ssRNA. Interestingly, the new atomic structures of the Csm proteins presented here are similar to those of previously known Csm proteins in other species despite their low sequence similarity. Our combined structural and biochemical data suggest that ssRNA cleavage is preferentially carried out near its 5'-end, that the extent of interactions among the ssRNA, crRNA and the protein components regulates the DNase activity of the Csm complex, and that the 3' flanking sequence of ssRNA activates the Cas10 DNase activity allosterically.
- HIT Center for Life Sciences, School of Life Science and Technology, Harbin Institute of Technology, Harbin, Heilongjiang, 150080, China.
Organizational Affiliation: