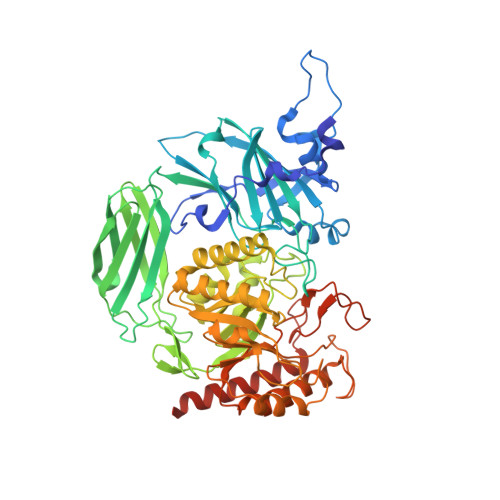
Selecting a Single Stereocenter: The Molecular Nuances That Differentiate beta-Hexuronidases in the Human Gut Microbiome.
Pellock, S.J., Walton, W.G., Redinbo, M.R.(2019) Biochemistry 58: 1311-1317
- PubMed: 30729778 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.8b01285
- Primary Citation Related Structures:
6NCW, 6NCX, 6NCY, 6NCZ - PubMed Abstract:
The human gut microbiome is a ripe space for the discovery of new proteins and novel functions. Many genes in the gut microbiome encode glycoside hydrolases that help bacteria scavenge sugars present in the human gut. Glycoside hydrolase family 2 (GH2) is one group of sugar-scavenging proteins, which includes β-glucuronidases (GUS) and β-galacturonidases (GalAses), enzymes that cleave the sugar conjugates of the epimers glucuronate and galacturonate. Here we structurally and functionally characterize a GH2 GalAse and a hybrid GUS/GalAse, which reveal the molecular details that enable these GHs to differentiate a single stereocenter. First, we characterized a previously annotated GUS from Eisenbergiella tayi and demonstrated that it is, in fact, a GalAse. We determined the crystal structure of this GalAse, identified the key residue that confers GalAse activity, and convert this GalAse into a GUS by mutating a single residue. We performed bioinformatic analysis of 279 putative GUS enzymes from the human gut microbiome and identified 12 additional putative GH2 GalAses, one of which we characterized and confirmed is a GalAse. Lastly, we report the structure of a hybrid GUS/GalAse from Fusicatenibacter saccharivorans, which revealed a unique hexamer that positions the N-terminus of adjacent protomers in the aglycone binding site. Taken together, these data reveal a new class of bacterial GalAses in the human gut microbiome and unravel the structural details that differentiate GH2 GUSs and GalAses.