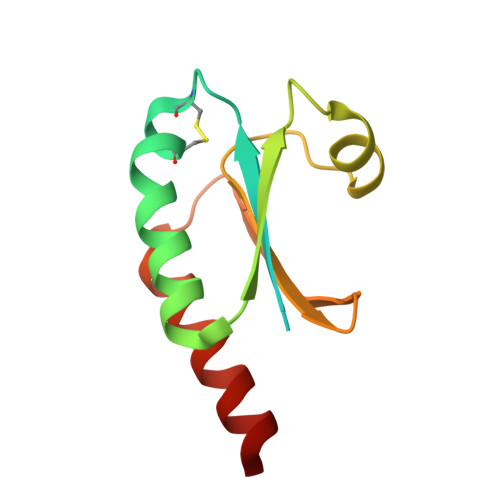
Structure and function of the putative thioredoxin 1 from the thermophilic eubacterium Thermosipho africanus strain TCF52B.
Sahtout, N., Kuttiyatveetil, J.R.A., Sanders, D.A.R.(2019) Biochim Biophys Acta Proteins Proteom 1867: 426-433
- PubMed: 30716506 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2019.01.011
- Primary Citation Related Structures:
6MOS - PubMed Abstract:
The thioredoxin system is a ubiquitous oxidoreductase system that consists of the enzyme thioredoxin reductase (TrxR), its cofactor nicotinamide adenine dinucleotide phosphate (NAD(P)H) and the protein thioredoxin (Trx). The system has been comprehensively studied from many organisms, such as Escherichia coli (E. coli); however, structural and functional analysis of this system from thermophilic bacteria has not been as extensive. In this study, Thermosipho africanus, a thermophilic eubacterium, Trx1 (TaTrx1) was successfully cloned, overexpressed and purified, to greater than 95% purity. Inspection of the amino acid sequence of TaTrx1 categorized the protein as a putative Trx. Its ability to reduce the interchain disulfides of insulin, in the presence of dithiothreitol, provided further evidence to suggest that it was a Trx. The three dimensional structure of the protein, determined using X-ray crystallography, provided additional evidence for this. The crystal structure was solved in space group P2 1 2 1 2 1 to 1.8 Ă resolution and showed the characteristic thioredoxin fold; four β-strands surrounded by three α-helices. The active site of TaTrx1 contained two cysteines that formed a disulfide bridge, and was structurally similar to the active site of EcTrx1. Further studies indicated that TaTrx1 was far more stable than Trx1 of E. coli (EcTrx1). The protein could withstand both higher temperatures and higher concentrations of guanidine hydrochloride before denaturing. Our studies have therefore identified a novel thermophilic putative Trx that structurally and functionally behaves like a Trx.
- Department of Chemistry, College of Arts and Science, University of Saskatchewan, Saskatoon, Saskatchewan, Canada.
Organizational Affiliation: