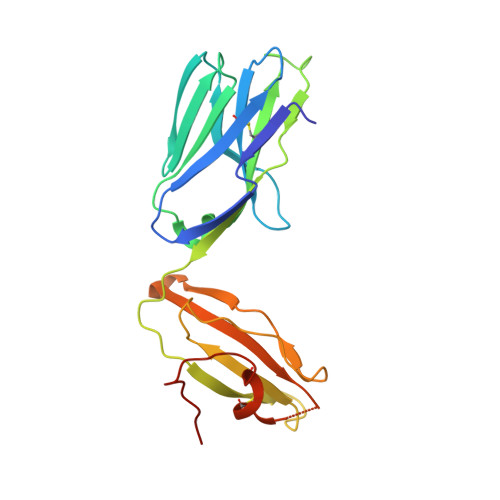
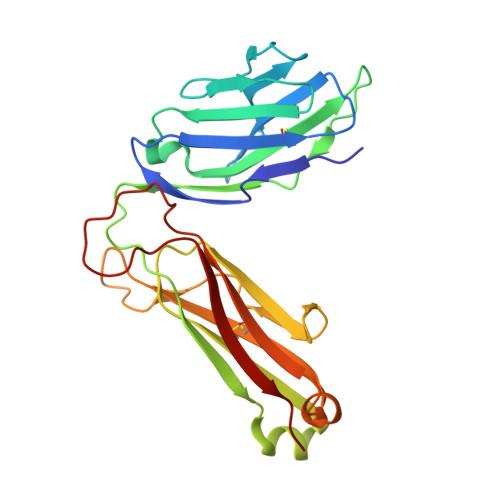
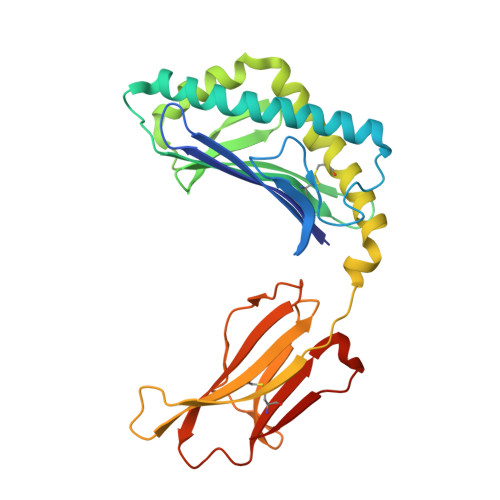
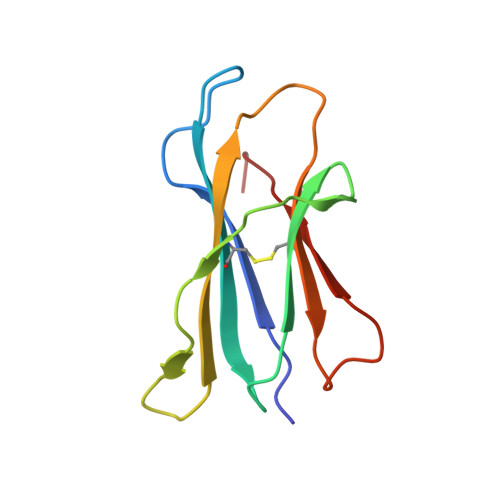
4"-O-Alkylated alpha-Galactosylceramide Analogues as iNKT-Cell Antigens: Synthetic, Biological, and Structural Studies.
Janssens, J., Bitra, A., Wang, J., Decruy, T., Venken, K., van der Eycken, J., Elewaut, D., Zajonc, D.M., van Calenbergh, S.(2019) ChemMedChem 14: 147-168
- PubMed: 30556652 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/cmdc.201800649
- Primary Citation Related Structures:
6MIV, 6MIY, 6MJ4, 6MJ6, 6MJA, 6MJI, 6MJJ, 6MJQ - PubMed Abstract:
Invariant natural killer T-cells (iNKT) are a glycolipid-responsive subset of T-lymphocytes that fulfill a pivotal role in the immune system. The archetypical synthetic glycolipid, α-galactosylceramide (α-GalCer), whose molecular framework is inspired by a group of amphiphilic natural products, remains the most studied antigen for iNKT-cells. Nonetheless, the potential of α-GalCer as an immunostimulating agent is compromised by the fact that this glycolipid elicits simultaneous secretion of Th1- and Th2-cytokines. This has incited medicinal chemistry efforts to identify analogues that are able to perturb the Th1/Th2 balance. In this work, we present the synthesis of an extensive set of 4"-O-alkylated α-GalCer analogues, which were evaluated in vivo for their cytokine induction. We have found that conversion of the 4"-OH group to ether moieties decreases the immunogenic potential in mice relative to α-GalCer. Yet, the benzyl-modified glycolipids are able to produce a distinct pro-inflammatory immune response. The crystal structures suggest an extra hydrophobic interaction between the benzyl moiety and the α2-helix of CD1d.
- Laboratory for Medicinal Chemistry, Department of Pharmaceutics (FFW), Ghent University, Ottergemsesteenweg 460, 9000, Ghent, Belgium.
Organizational Affiliation: