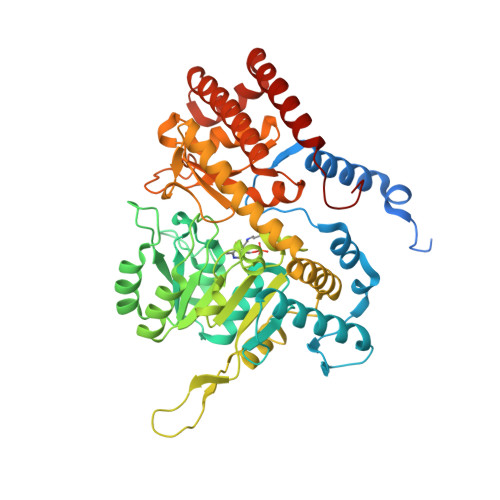
Structural basis for selective inhibition of human serine hydroxymethyltransferase by secondary bile acid conjugate.
Ota, T., Senoo, A., Shirakawa, M., Nonaka, H., Saito, Y., Ito, S., Ueno, G., Nagatoishi, S., Tsumoto, K., Sando, S.(2021) iScience 24: 102036-102036
- PubMed: 33521601 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.isci.2021.102036
- Primary Citation Related Structures:
6M5O, 6M5W - PubMed Abstract:
Bile acids are metabolites of cholesterol that facilitate lipid digestion and absorption in the small bowel. Bile acids work as agonists of receptors to regulate their own metabolism. Bile acids also regulate other biological systems such as sugar metabolism, intestinal multidrug resistance, and adaptive immunity. However, numerous physiological roles of bile acids remain undetermined. In this study, we solved the crystal structure of human serine hydroxymethyltransferase (hSHMT) in complex with an endogenous secondary bile acid glycine conjugate. The specific interaction between hSHMT and the ligand was demonstrated using mutational analyses, biophysical measurements, and structure-activity relationship studies, suggesting that secondary bile acid conjugates may act as modulators of SHMT activity.
- Department of Chemistry and Biotechnology, Graduate School of Engineering, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo, 113-8656, Japan.
Organizational Affiliation: