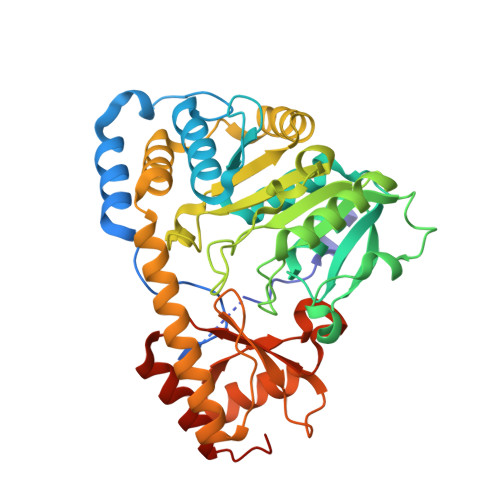
Structural insights into the catalytic mechanism of Bacillus subtilis BacF.
Deshmukh, A., Gopal, B.(2020) Acta Crystallogr F Struct Biol Commun 76: 145-151
- PubMed: 32134000 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X20001636
- Primary Citation Related Structures:
6L1L, 6L1N, 6L1O - PubMed Abstract:
The nonribosomal biosynthesis of the dipeptide antibiotic bacilysin is achieved by the concerted action of multiple enzymes in the Bacillus subtilis bac operon. BacF (YwfG), encoded by the bacF gene, is a fold type I pyridoxal 5-phosphate (PLP)-dependent stereospecific transaminase. Activity assays with L-phenylalanine and 4-hydroxyphenylpyruvic acid (4HPP), a chemical analogue of tetrahydrohydroxyphenylpyruvic acid (H 4 HPP), revealed stereospecific substrate preferences, a finding that was consistent with previous reports on the role of this enzyme in bacilysin synthesis. The crystal structure of this dimeric enzyme was determined in its apo form as well as in substrate-bound and product-bound conformations. Two ligand-bound structures were determined by soaking BacF crystals with substrates (L-phenylalanine and 4-hydroxyphenylpyruvate). These structures reveal multiple catalytic steps: the internal aldimine with PLP and two external aldimine conformations that show the rearrangement of the external aldimine to generate product (L-tyrosine). Together, these structural snapshots provide an insight into the catalytic mechanism of this transaminase.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore, Karnataka 560 012, India.
Organizational Affiliation: