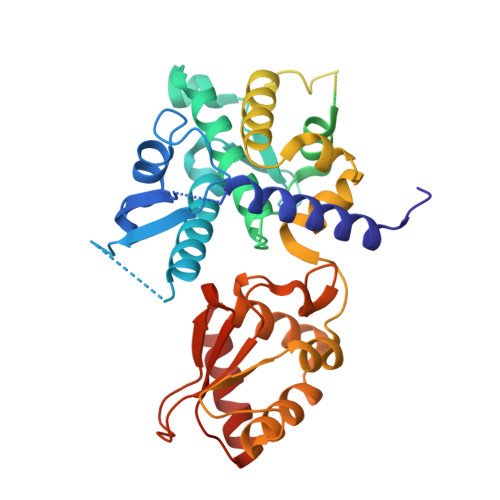
Crystal structure of HECT domain of UBE3C E3 ligase and its ubiquitination activity.
Singh, S., Sivaraman, J.(2020) Biochem J 477: 905-923
- PubMed: 32039437
- DOI: https://doi.org/10.1042/BCJ20200027
- Primary Citation Related Structures:
6K2C - PubMed Abstract:
The HECT family of E3 ubiquitin ligase is divided into three subfamilies: the NEDD4, the HERC, and the 'other'. Previous studies have mostly targeted members of the NEDD4 subfamily for structural and functional analysis. The UBE3C E3 ligase is a member of the 'other' subfamily HECT and influences several crucial cellular processes, including innate immunity, proteasome processivity, and cancer metastasis. Here, we report the crystal structure of the HECT domain of UBE3C (amino acids (aa) 744-1083) with an additional fifty N-terminal amino acids (aa 693-743) at 2.7 Å, along with multiple in vitro ubiquitination assays to understand its enzymatic activity. The UBE3C HECT domain forms an open, L-shaped, bilobed conformation, having a large N-lobe and a small C-lobe. We show that the N-terminal region (aa 693-743) preceding the UBE3C HECT domain as well as a loop region (aa 758-762) in the N-lobe of the HECT domain affect the stability and activity of UBE3C HECT domain. Moreover, we identified Lys903 in the UBE3C HECT domain as a major site of autoubiquitination. The deletion of the last three amino acids at the C-terminal completely abrogated UBE3C activity while mutations of Gln961 and Ser1049 residues in the HECT domain substantially decreased its autoubiquitination activity. We demonstrate that these region/residues are involved in the E2-E3 transthiolation process and affect the UBE3C mediated autoubiquitination. Collectively, our study identified key residues crucial for UBE3C enzymatic activity, and it may assist in the development of suitable inhibitors to regulate its activity in multiple cancers.
- Department of Biological Sciences, National University of Singapore, 14 Science Drive 4, Singapore 117543, Singapore.
Organizational Affiliation: