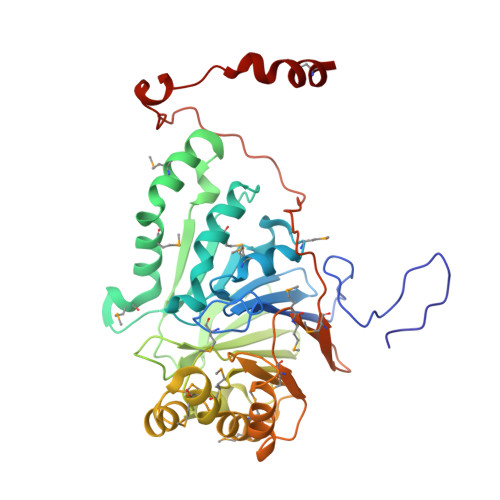
PNGM 1 a novel subclass B3 metallo beta lactamase from a deep sea sediment metagenome
Kang, L.W., Lee, S.H.(2018) J Glob Antimicrob Resist 14: 302-305
- PubMed: 29842976 Search on PubMed
- DOI: https://doi.org/10.1016/j.jgar.2018.05.021
- Primary Citation Related Structures:
6JKW - PubMed Abstract:
In order to find antimicrobial resistance gene(s) pre-dating the use of antibiotics through metagenomics, functional screening of a metagenomic library from the deep-seep sediments of Edison Seamount (ca. 10000 years old) was performed. Among 60 antimicrobial-resistant clones, a single clone with the highest minimum inhibitory concentration (MIC) for ampicillin was selected. Sequence analysis revealed a new metallo-β-lactamase (MBL) gene, designated as bla PNGM-1 . PNGM-1 retains a zinc ion-binding motif (H 116 XH 118 XD 120 H 121 , H 196 and H 263 ), conserved in subclass B3 MBLs. The catalytic parameters of purified PNGM-1 and the MICs of β-lactams for Escherichia coli TOP10 transformants harbouring the bla PNGM-1 gene were assessed. Antimicrobial susceptibility testing indicated reduced susceptibility to penicillins, narrow- and extended-spectrum cephalosporins, and carbapenems in E. coli TOP10 transformants harbouring the bla PNGM-1 gene. In addition, kinetic analyses revealed that PNGM-1 hydrolysed almost all β-lactams. The PNGM-1 enzyme is the first case of a subclass B3 MBL derived from a functional metagenomic library of a deep-sea sediment that pre-dates the antibiotic era.
- National Leading Research Laboratory of Drug Resistance Proteomics, Department of Biological Sciences, Myongji University, 116 Myongjiro, Yongin, Gyeonggido 17058, Republic of Korea.
Organizational Affiliation: