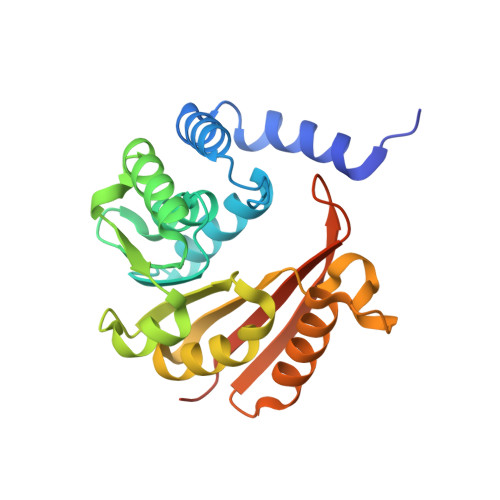
Structural and biochemical characterization of Rv0187, an O-methyltransferase from Mycobacterium tuberculosis.
Lee, S., Kang, J., Kim, J.(2019) Sci Rep 9: 8059-8059
- PubMed: 31147608 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-44592-7
- Primary Citation Related Structures:
6JCL, 6JCM - PubMed Abstract:
Catechol O-methyltransferase (COMT) is widely distributed in nature and installs a methyl group onto one of the vicinal hydroxyl groups of a catechol derivative. Enzymes belonging to this family require two cofactors for methyl transfer: S-adenosyl-l-methionine as a methyl donor and a divalent metal cation for regiospecific binding and activation of a substrate. We have determined two high-resolution crystal structures of Rv0187, one of three COMT paralogs from Mycobacterium tuberculosis, in the presence and absence of cofactors. The cofactor-bound structure clearly locates strontium ions and S-adenosyl-l-homocysteine in the active site, and together with the complementary structure of the ligand-free form, it suggests conformational dynamics induced by the binding of cofactors. Examination of in vitro activities revealed promiscuous substrate specificity and relaxed regioselectivity against various catechol-like compounds. Unexpectedly, mutation of the proposed catalytic lysine residue did not abolish activity but altered the overall landscape of regiospecific methylation.
- Department of Chemistry, Gwangju Institute of Science and Technology, Gwangju, 61005, Republic of Korea.
Organizational Affiliation: