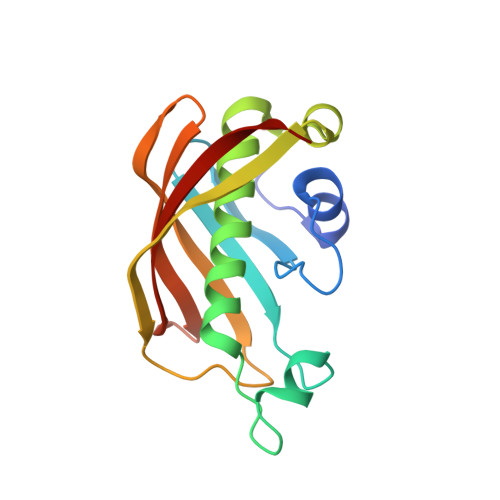
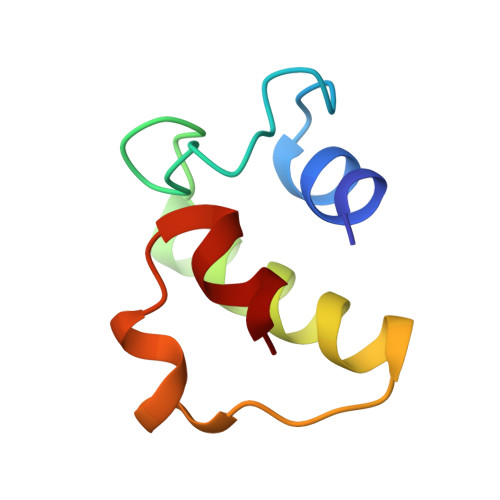
A back-door Phenylalanine coordinates the stepwise hexameric loading of acyl carrier protein by the fatty acid biosynthesis enzyme beta-hydroxyacyl-acyl carrier protein dehydratase (FabZ).
Shen, S.Q., Hang, X.D., Zhuang, J.J., Zhang, L., Bi, H.K., Zhang, L.(2019) Int J Biol Macromol 128: 5-11
- PubMed: 30677439
- DOI: https://doi.org/10.1016/j.ijbiomac.2019.01.094
- Primary Citation Related Structures:
6IHC - PubMed Abstract:
The fatty acid biosynthesis pathway (FAS) was a fundamental procedure to generate a diversity of lipid components for cellular metabolism in bacteria, while the mechanism of substrate recognition remains unclear. The β-hydroxyacyl-acyl carrier protein dehydratase hexamer (FabZ) is an essential module in the elongation cycle of type-II FAS, catalyzing the dehydration of β-hydroxyacyl-lipid substrate carried by the holo form acyl carrier protein (holo-ACP). We previously elucidated an alternating seesaw-like ACP loading manner within a FabZ dimer subunits, mediated by a front-door residue Tyrosine (Tyr100). Here, we demonstrated that a back-door residue Phenylalanine (Phe83) of FabZ regulates the stepwise hexameric loading of ACP. Our finding represents clues as to the dynamic ACP recognition and catalysis mechanism of dehydratase in fatty acid biosynthesis, and provides critical information for developing antimicrobials targeting the dehydratase module in fatty acid biosynthesis pathway.
- Department of Pharmacology and Chemical Biology, Shanghai Jiao Tong University School of Medicine, Shanghai, PR China; Shanghai Universities Collaborative Innovation Center for Translational Medicine, Shanghai, PR China.
Organizational Affiliation: