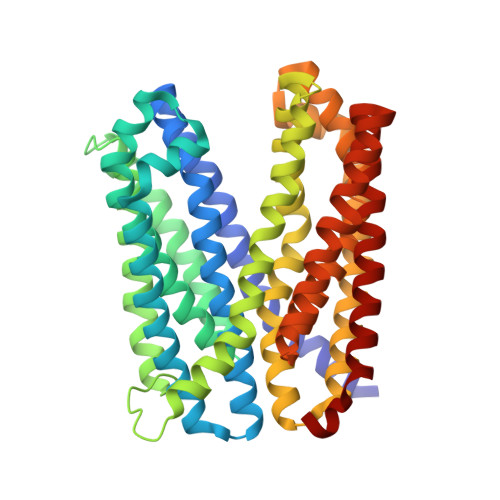
Structural Basis of H+-Dependent Conformational Change in a Bacterial MATE Transporter.
Kusakizako, T., Claxton, D.P., Tanaka, Y., Maturana, A.D., Kuroda, T., Ishitani, R., Mchaourab, H.S., Nureki, O.(2019) Structure 27: 293
- PubMed: 30449688 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2018.10.004
- Primary Citation Related Structures:
6IDP, 6IDR, 6IDS - PubMed Abstract:
Multidrug and toxic compound extrusion (MATE) transporters efflux toxic compounds using a Na + or H + gradient across the membrane. Although the structures of MATE transporters have been reported, the cation-coupled substrate transport mechanism remains controversial. Here we report crystal structures of VcmN, a Vibrio cholerae MATE transporter driven by the H + gradient. High-resolution structures in two distinct conformations associated with different pHs revealed that the rearrangement of the hydrogen-bonding network around the conserved Asp35 induces the bending of transmembrane helix 1, as in the case of the H + -coupled Pyrococcus furiosus MATE transporter. We also determined the crystal structure of the D35N mutant, which captured a unique conformation of TM1 facilitated by an altered hydrogen-bonding network. Based on the present results, we propose a common step in the transport cycle shared among prokaryotic H + -coupled MATE transporters.
- Department of Biological Sciences, Graduate School of Science, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: