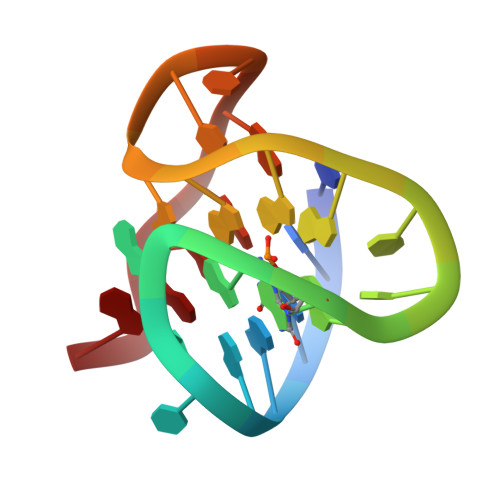
Impact of Oxidative Lesions on the Human Telomeric G-Quadruplex.
Bielskute, S., Plavec, J., Podbevsek, P.(2019) J Am Chem Soc 141: 2594-2603
- PubMed: 30657306 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.8b12748
- Primary Citation Related Structures:
6IA0, 6IA4 - PubMed Abstract:
Telomere attrition is closely associated with cell aging and exposure to reactive oxygen species (ROS). While oxidation products of nucleotides have been studied extensively in the past, the underlying secondary/tertiary structural changes in DNA remain poorly understood. In this work, we systematically probed guanine positions in the human telomeric oligonucleotide sequence (hTel) by substitutions with the major product of ROS, 8-oxo-7,8-dihydroguanine ( oxo G), and evaluated the G-quadruplex forming ability of such oligonucleotides. Due to reduced hydrogen-bonding capability caused by oxo G, a loss of G-quadruplex structure was observed for most oligonucleotides containing oxidative lesions. However, some positions in the hTel sequence were found to tolerate substitutions with oxo G. Due to oxo G's preference for the syn conformation, distinct responses were observed when replacing guanines with different glycosidic conformations. Accommodation of oxo G at sites originally in syn or anti in nonsubstituted hTel G-quadruplex requires a minor structural rearrangement or a major conformational shift, respectively. The system responds by retaining or switching to a fold where oxo G is in syn conformation. Most importantly, these G-quadruplex structures are still stable at physiological temperatures and should be considered detrimental in higher-order telomere structures.
- Slovenian NMR Center , National Institute of Chemistry , Hajdrihova 19 , SI-1000 Ljubljana , Slovenia.
Organizational Affiliation: