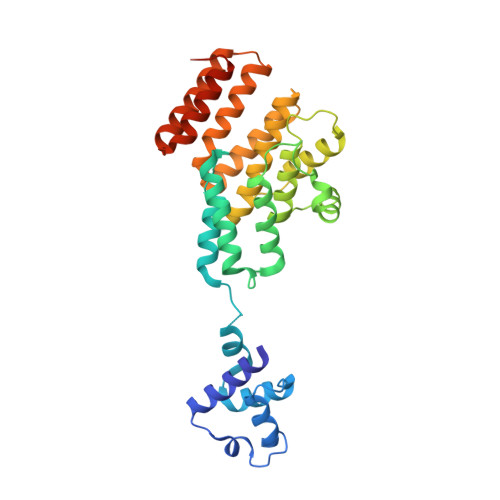
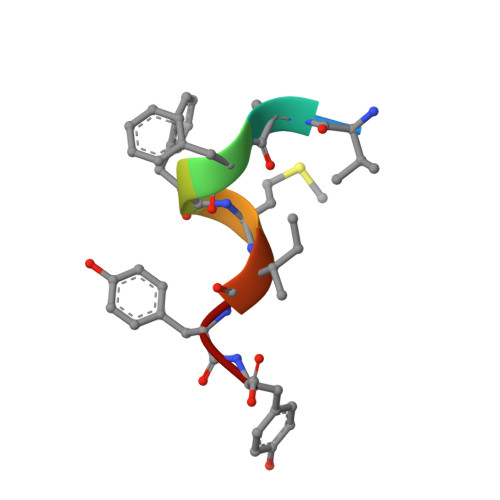
Molecular dissection of pheromone selectivity in the competence signaling system ComRS of streptococci.
Ledesma-Garcia, L., Thuillier, J., Guzman-Espinola, A., Ensinck, I., Li de la Sierra-Gallay, I., Lazar, N., Aumont-Nicaise, M., Mignolet, J., Soumillion, P., Nessler, S., Hols, P.(2020) Proc Natl Acad Sci U S A 117: 7745-7754
- PubMed: 32198205
- DOI: https://doi.org/10.1073/pnas.1916085117
- Primary Citation of Related Structures:
6HU8, 6HUA, 6QER - PubMed Abstract:
Competence allows bacteria to internalize exogenous DNA fragments for the acquisition of new phenotypes such as antibiotic resistance or virulence traits. In most streptococci, competence is regulated by ComRS signaling, a system based on the mature ComS pheromone (XIP), which is internalized to activate the (R)RNPP-type ComR sensor by triggering dimerization and DNA binding. Cross-talk analyses demonstrated major differences of selectivity between ComRS systems and raised questions concerning the mechanism of pheromone-sensor recognition and coevolution. Here, we decipher the molecular determinants of selectivity of the closely related ComRS systems from Streptococcus thermophilus and Streptococcus vestibularis Despite high similarity, we show that the divergence in ComR-XIP interaction does not allow reciprocal activation. We perform the structural analysis of the ComRS system from S. vestibularis. Comparison with its ortholog from S. thermophilus reveals an activation mechanism based on a toggle switch involving the recruitment of a key loop by the XIP C terminus. Together with a broad mutational analysis, we identify essential residues directly involved in peptide binding. Notably, we generate a ComR mutant that displays a fully reversed selectivity toward the heterologous pheromone with only five point mutations, as well as other ComR variants featuring XIP bispecificity and/or neofunctionalization for hybrid XIP peptides. We also reveal that a single XIP mutation relaxes the strictness of ComR activation, suggesting fast adaptability of molecular communication phenotypes. Overall, this study is paving the way toward the rational design or directed evolution of artificial ComRS systems for a range of biotechnological and biomedical applications.
- Louvain Institute of Biomolecular Science and Technology, Biochemistry and Genetics of Microorganisms, Université catholique de Louvain, B-1348 Louvain-La-Neuve, Belgium.
Organizational Affiliation: