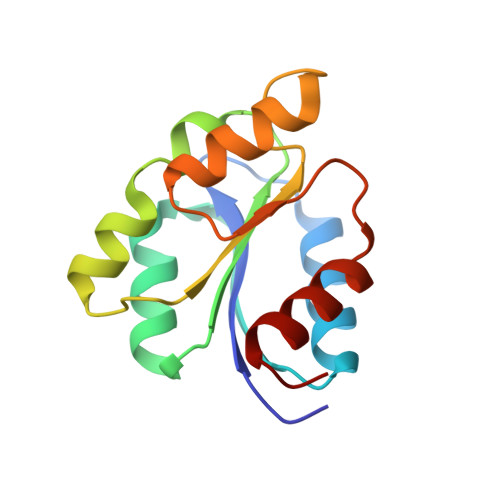
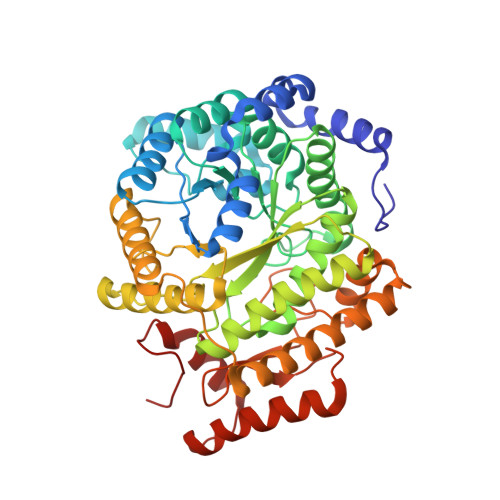
Structure-Based Demystification of Radical Catalysis by a Coenzyme B 12 Dependent Enzyme-Crystallographic Study of Glutamate Mutase with Cofactor Homologues.
Gruber, K., Csitkovits, V., Lyskowski, A., Kratky, C., Krautler, B.(2022) Angew Chem Int Ed Engl 61: e202208295-e202208295
- PubMed: 35793207 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/anie.202208295
- Primary Citation Related Structures:
6H9E, 6H9F - PubMed Abstract:
Catalysis by radical enzymes dependent on coenzyme B 12 (AdoCbl) relies on the reactive primary 5'-deoxy-5'adenosyl radical, which originates from reversible Co-C bond homolysis of AdoCbl. This bond homolysis is accelerated roughly 10 12 -fold upon binding the enzyme substrate. The structural basis for this activation is still strikingly enigmatic. As revealed here, a displaced firm adenosine binding cavity in substrate-loaded glutamate mutase (GM) causes a structural misfit for intact AdoCbl that is relieved by the homolytic Co-C bond cleavage. Strategically interacting adjacent adenosine- and substrate-binding protein cavities provide a tight caged radical reaction space, controlling the entire radical path. The GM active site is perfectly structured for promoting radical catalysis, including "negative catalysis", a paradigm for AdoCbl-dependent mutases.
- Institute of Molecular Biosciences, University of Graz, Humboldtstraße 50, 8010, Graz, Austria.
Organizational Affiliation: