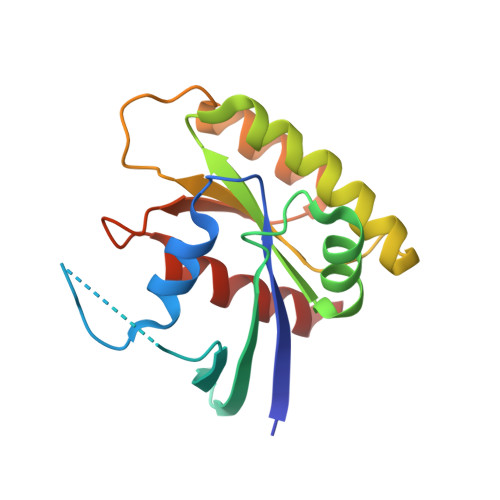
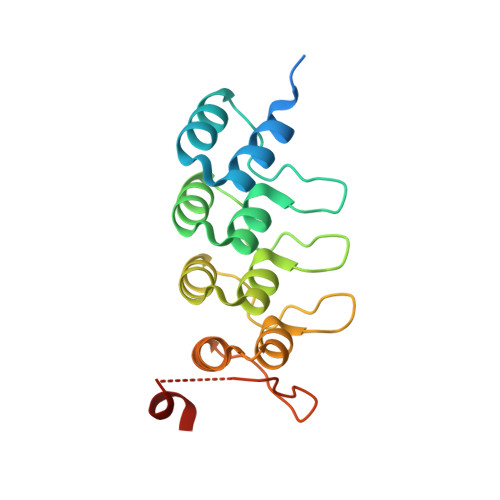
KRAS-specific inhibition using a DARPin binding to a site in the allosteric lobe.
Bery, N., Legg, S., Debreczeni, J., Breed, J., Embrey, K., Stubbs, C., Kolasinska-Zwierz, P., Barrett, N., Marwood, R., Watson, J., Tart, J., Overman, R., Miller, A., Phillips, C., Minter, R., Rabbitts, T.H.(2019) Nat Commun 10: 2607-2607
- PubMed: 31197133 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-10419-2
- Primary Citation Related Structures:
6H46, 6H47 - PubMed Abstract:
Inhibiting the RAS oncogenic protein has largely been through targeting the switch regions that interact with signalling effector proteins. Here, we report designed ankyrin repeat proteins (DARPins) macromolecules that specifically inhibit the KRAS isoform by binding to an allosteric site encompassing the region around KRAS-specific residue histidine 95 at the helix α3/loop 7/helix α4 interface. We show that these DARPins specifically inhibit KRAS/effector interactions and the dependent downstream signalling pathways in cancer cells. Binding by the DARPins at that region influences KRAS/effector interactions in different ways, including KRAS nucleotide exchange and inhibiting KRAS dimerization at the plasma membrane. These results highlight the importance of targeting the α3/loop 7/α4 interface, a previously untargeted site in RAS, for specifically inhibiting KRAS function.
- Weatherall Institute of Molecular Medicine, MRC Molecular Haematology Unit, University of Oxford, John Radcliffe Hospital, Oxford, OX3 9DS, UK.
Organizational Affiliation: