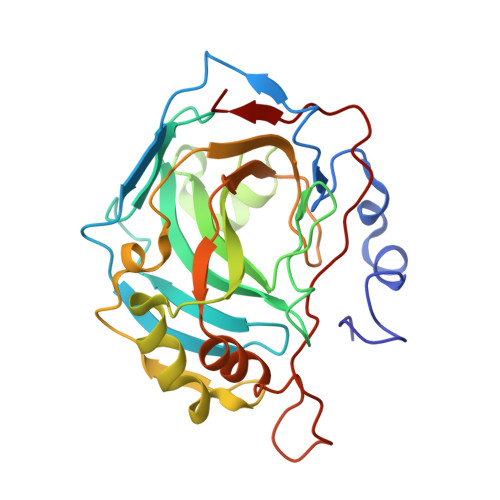
Inhibitor-Polymer Conjugates as a Versatile Tool for Detection and Visualization of Cancer-Associated Carbonic Anhydrase Isoforms
Pospisilova, K., Knedlik, T., Sacha, P., Kostka, L., Schimer, J., Brynda, J., Kral, V., Cigler, P., Navratil, V., Etrych, T., Subr, V., Kugler, M., Fabry, M., Rezacova, P., Konvalinka, J.(2019) ACS Omega