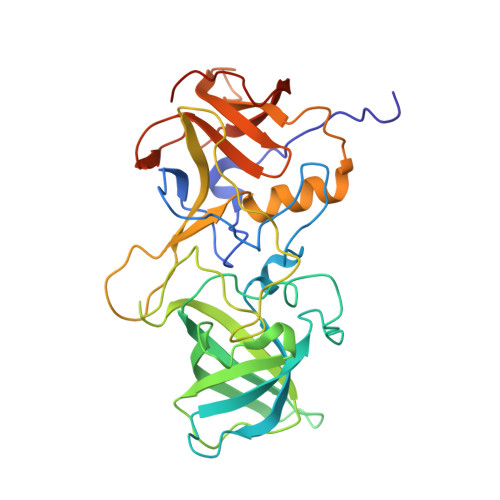
Structural Basis for Human Norovirus Capsid Binding to Bile Acids.
Kilic, T., Koromyslova, A., Hansman, G.S.(2019) J Virol 93
- PubMed: 30355683
- DOI: https://doi.org/10.1128/JVI.01581-18
- Primary Citation Related Structures:
6GVZ, 6GW0, 6GW1, 6GW2, 6GW4 - PubMed Abstract:
A recently developed human norovirus cell culture system revealed that the presence of bile enhanced or was an essential requirement for the growth of certain genotypes. Before this discovery, histo-blood group antigens (HBGAs) were the only well-studied cofactor known for human noroviruses, and there was evidence that several genotypes poorly bound HBGAs. Therefore, the purpose of this study was to investigate how human norovirus capsids interact with bile acids. We found that bile acids had low-micromolar affinities for GII.1, GII.10, and GII.19 capsids but did not bind GI.1, GII.3, GII.4, or GII.17. We showed that bile acid bound at a partially conserved pocket on the norovirus capsid-protruding (P) domain using X-ray crystallography. Amino acid sequence alignment and structural analysis delivered an explanation of selective bile acid binding. Intriguingly, we discovered that binding of the bile acid was the critical step to stabilize several P domain loops that optimally placed an essential amino acid side chain (Asp375) to bind HBGAs in an otherwise HBGA nonbinder (GII.1). Furthermore, bile acid enhanced HBGA binding for a known HBGA binder (GII.10). Altogether, these new data suggest that bile acid functions as a loop-stabilizing regulator and enhancer of HBGA binding for certain norovirus genotypes. IMPORTANCE Given that human norovirus virions likely interact with bile acid during a natural infection, our evidence that an HBGA nonbinder (GII.1) can be converted to an HBGA binder after bile acid binding is of major significance. Our data provide direct evidence that, like HBGAs, bile acid interaction on the capsid is an important cofactor for certain genotypes. However, more unanswered questions seem to arise from these new discoveries. For example, is there an association between the bile acid requirement and the prevalence of certain genotypes? That is, the GII.1 and GII.10 (bile acid binders) genotypes rarely caused outbreaks, whereas the GII.4 and GII.17 genotypes (bile acid nonbinders) were responsible for large epidemics. Therefore, it seems plausible that certain genotypes require bile acids, whereas others have modified their bile acid requirements on the capsid.
- Schaller Research Group at the University of Heidelberg and the German Cancer Research Center, Heidelberg, Germany.
Organizational Affiliation: