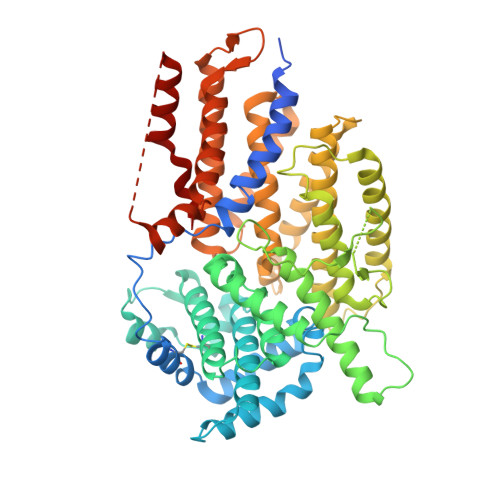
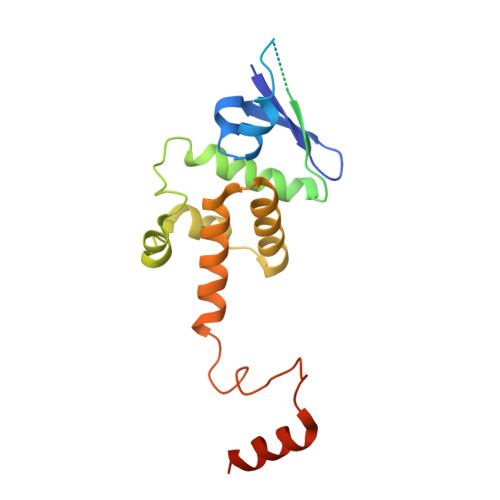
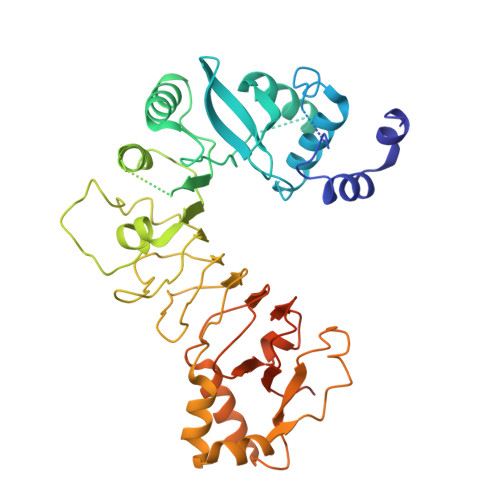
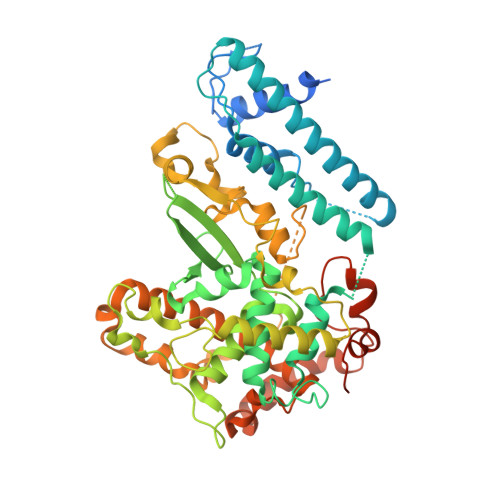
Insights into Centromere DNA Bending Revealed by the Cryo-EM Structure of the Core Centromere Binding Factor 3 with Ndc10.
Zhang, W., Lukoynova, N., Miah, S., Lucas, J., Vaughan, C.K.(2018) Cell Rep 24: 744-754
- PubMed: 30021170 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2018.06.068
- Primary Citation Related Structures:
6FE8, 6GSA - PubMed Abstract:
The centromere binding factor 3 (CBF3) complex binds the third centromere DNA element in organisms with point centromeres, such as S. cerevisiae. It is an essential complex for assembly of the kinetochore in these organisms, as it facilitates genetic centromere specification and allows association of all other kinetochore components. We determined high-resolution structures of the core complex of CBF3 alone and in association with a monomeric construct of Ndc10, using cryoelectron microscopy (cryo-EM). We identify the DNA-binding site of the complex and present a model in which CBF3 induces a tight bend in centromeric DNA, thus facilitating assembly of the centromeric nucleosome.
- Institute of Structural and Molecular Biology, Birkbeck College, Malet Street, London WC1E 7HX, UK; Institute of Structural and Molecular Biology, University College London, Gower Street, London WC1E 6BT, UK.
Organizational Affiliation: