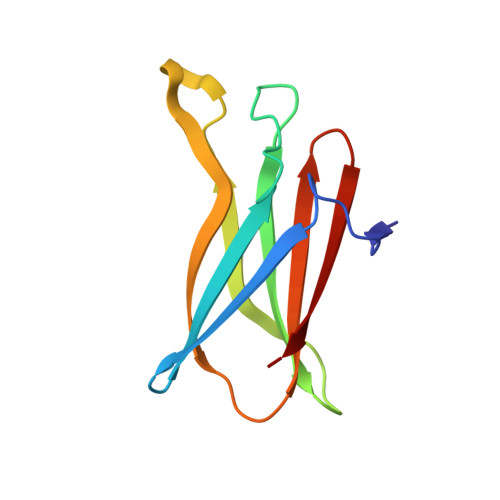
Insights into the Structures of Superoxide Reductases from the Symbionts Ignicoccus hospitalis and Nanoarchaeum equitans.
Romao, C.V., Matias, P.M., Sousa, C.M., Pinho, F.G., Pinto, A.F., Teixeira, M., Bandeiras, T.M.(2018) Biochemistry 57: 5271-5281
- PubMed: 29939726
- DOI: https://doi.org/10.1021/acs.biochem.8b00334
- Primary Citation of Related Structures:
6GQ8 - PubMed Abstract:
Superoxide reductases (SORs) are enzymes that detoxify the superoxide anion through its reduction to hydrogen peroxide and exist in both prokaryotes and eukaryotes. The substrate is transformed at an iron catalytic center, pentacoordinated in the ferrous state by four histidines and one cysteine. SORs have a highly conserved motif, (E)(K)HxP-, in which the glutamate is associated with a redox-driven structural change, completing the octahedral coordination of the iron in the ferric state, whereas the lysine may be responsible for stabilization and donation of a proton to catalytic intermediates. We aimed to understand at the structural level the role of these two residues, by determining the X-ray structures of the SORs from the hyperthermophilic archaea Ignicoccus hospitalis and Nanoarchaeum equitans that lack the quasi-conserved lysine and glutamate, respectively, but have catalytic rate constants similar to those of the canonical enzymes, as we previously demonstrated. Furthermore, we have determined the crystal structure of the E23A mutant of I. hospitalis SOR, which mimics several enzymes that lack both residues. The structures revealed distinct structural arrangements of the catalytic center that simulate several catalytic cycle intermediates, namely, the reduced and the oxidized forms, and the glutamate-free and deprotonated ferric forms. Moreover, the structure of the I. hospitalis SOR provides evidence for the presence of an alternative lysine close to the iron center in the reduced state that may be a functional substitute for the "canonical" lysine.
- ITQB NOVA, Instituto de Tecnologia Química e Biológica António Xavier , Universidade Nova de Lisboa , Av. da República , 2780-157 Oeiras , Portugal.
Organizational Affiliation: