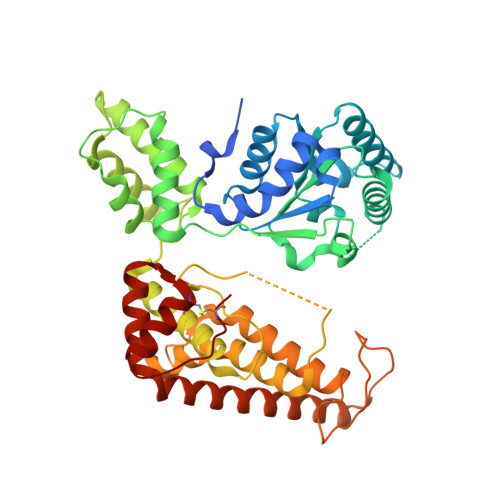
Conformational flexibility of pore loop-1 gives insights into substrate translocation by the AAA+protease FtsH.
Uthoff, M., Baumann, U.(2018) J Struct Biol 204: 199-206
- PubMed: 30118817
- DOI: https://doi.org/10.1016/j.jsb.2018.08.009
- Primary Citation Related Structures:
6GCN, 6GCO - PubMed Abstract:
Two crystal structures of a transmembrane helix-lacking FtsH construct from Aquifex aeolicus have been determined at 2.9 Å and 3.3 Å resolution in space groups R32 and P312, respectively. Both structures are virtually identical despite different crystal packing contacts. In both structures, the FtsH hexamer is created from two different subunits of the asymmetric unit by the threefold symmetry of the crystals. Similar to other published structures, all subunits are loaded with ADP and the two subunit in the asymmetric unit resemble the already known open and closed conformations. Within the ATPase cycle while the whole subunit switches from the opened to the closed state, pore loop-1 interacts with the substrate and translocates it into the proteolytic chamber. Unique to our models is a presumably inactive conformation of the pore loop which allows the closed conformation to switch back to the opened state without pushing the substrate out again. Our structures give further insights on how this new pore loop conformation is induced and how it is linked to the intersubunit signalling network.
- Institute of Biochemistry, University of Cologne, Zuelpicher Str. 47, Cologne, North Rhine-Westphalia 50674, Germany.
Organizational Affiliation: