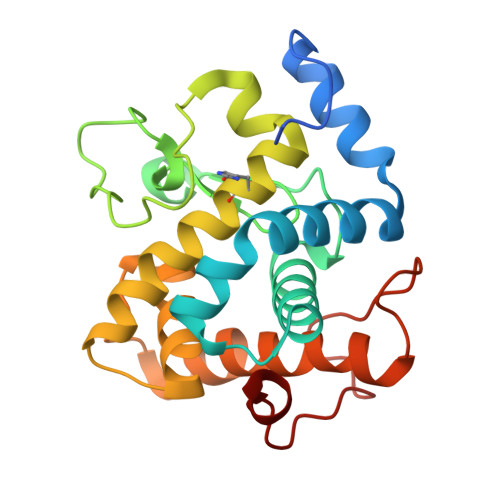
Structural studies of the unusual metal-ion site of the GH124 endoglucanase from Ruminiclostridium thermocellum.
Urresti, S., Cartmell, A., Liu, F., Walton, P.H., Davies, G.J.(2018) Acta Crystallogr F Struct Biol Commun 74: 496-505
- PubMed: 30084399
- DOI: https://doi.org/10.1107/S2053230X18006842
- Primary Citation of Related Structures:
6G1G, 6G1I - PubMed Abstract:
The recent discovery of `lytic' polysaccharide monooxygenases, copper-dependent enzymes for biomass degradation, has provided new impetus for the analysis of unusual metal-ion sites in carbohydrate-active enzymes. In this context, the CAZY family GH124 endoglucanase from Ruminiclostridium thermocellum contains an unusual metal-ion site, which was originally modelled as a Ca 2+ site but features aspartic acid, asparagine and two histidine imidazoles as coordinating residues, which are more consistent with a transition-metal binding environment. It was sought to analyse whether the GH124 metal-ion site might accommodate other metals. It is demonstrated through thermal unfolding experiments that this metal-ion site can accommodate a range of transition metals (Fe 2+ , Cu 2+ , Mn 2+ and Ni 2+ ), whilst the three-dimensional structure and mass spectrometry show that one of the histidines is partially covalently modified and is present as a 2-oxohistidine residue; a feature that is rarely observed but that is believed to be involved in an `off-switch' to transition-metal binding. Atomic resolution (<1.1 Å) complexes define the metal-ion site and also reveal the binding of an unusual fructosylated oligosaccharide, which was presumably present as a contaminant in the cellohexaose used for crystallization. Although it has not been possible to detect a biological role for the unusual metal-ion site, this work highlights the need to study some of the many metal-ion sites in carbohydrate-active enzymes that have long been overlooked or previously mis-assigned.
- York Structural Biology Laboratory, Department of Chemistry, University of York, York YO10 5DD, England.
Organizational Affiliation: